Cancer Center
 The CSHL Cancer Center is a basic research facility committed to exploring the fundamental biology of human cancer. With support from the National Cancer Institute (NCI) since 1987, our researchers have used a focused, multi-disciplinary approach to break new ground in basic tumor biology and develop innovative, advanced technologies. Research covers a broad range of cancer types, including breast, prostate, leukemia, glioma, pancreatic, sarcoma, lung, and melanoma.
The CSHL Cancer Center is a basic research facility committed to exploring the fundamental biology of human cancer. With support from the National Cancer Institute (NCI) since 1987, our researchers have used a focused, multi-disciplinary approach to break new ground in basic tumor biology and develop innovative, advanced technologies. Research covers a broad range of cancer types, including breast, prostate, leukemia, glioma, pancreatic, sarcoma, lung, and melanoma.
Three Scientific Programs provide focus in Cancer Genetics and Genomics; Cellular Communication in Cancer; and Gene Regulation and Inheritance. In addition, ten Shared Resources offer essential access to technologies, services, and expertise that enhance productivity. With a strong collaborative environment and open communication, the CSHL Cancer Center is able to make breakthroughs in cancer biology that are translating into real progress in cancer diagnostics and treatment.
Members of the CSHL Cancer Center apply a multi-pronged approach—from genomic biology to animal models to detailed biochemistry—to interrogate the molecular mechanisms that drive tumor growth and metastasis. Building on this basic research, scientists at the Lab are translating their findings into novel therapeutics for many of the most intractable cancers. Much of this research is made possible through numerous collaborations with clinical partners, including a strategic alliance with the nearby Northwell Health System that connects CSHL scientists with clinicians and more than 16,000 cancer patients each year.
Leadership and Administration
Leadership
Director: David Tuveson, M.D., Ph.D.
Deputy Director: Chris Vakoc, M.D., Ph.D.
Associate Director, Administration: Lindsey Baker, Ph.D.
Associate Director, Shared Resources: Nicholas Tonks, Ph.D.
Associate Director, Education, Lloyd Trotman, Ph.D.
Assistant Director, Shared Resources: Michael Lukey, Ph.D.
Administration
Assistant Director of Administration, Research: Katie Brenner
Assistant Director of Administration, Education and Leadership: Jessica Peluso
Director, Clinical & Translational Collaborations at CSHL: Soma Prum
Assistant Director, Shared Resources Operations: Denise Roberts, Ph.D.
Coordinator: Kelly Lewis
Program leaders
Co-program leaders, Cancer Genetics and Genomics: Tobias Janowitz, M.D., Ph.D., Camila dos Santos, Ph.D.
Co-program leaders, Cellular Communication in Cancer: Corina Amor Vegas, M.D., Ph.D., Linda Van Aelst, Ph.D.
Co-program leaders, Gene Regulation and Inheritance: Justin Kinney, Ph.D., Adrian Krainer, Ph.D.
Cancer Center External Advisory Board
Marcia Cruz, M.D., Ph.D.
Director, Gastrointestinal Oncology Division, University of Puerto Rico Comprehensive Cancer Center
Michael Kastan, M.D., Ph.D.
Executive Director, Duke Cancer Center
Wendy Law, Ph.D. (Administrative Advisor)
Associate Director of Administration, Fred Hutch/University of Washington/Seattle Children’s Cancer Consortium
Elaine Mardis, Ph.D.
Co-Executive Director, The Institute for Genomic Medicine at Nationwide Children’s Hospital and the Nationwide Foundation Endowed Chair of Genomic Medicine
Larry Norton, M.D.
Deputy Physician-in-Chief, Breast Cancer Programs
Medical Director, Evelyn H. Lauder Breast Center, Memorial Sloan Kettering Cancer Center
Dana Pe’er, Ph.D.
Scientific Director, The Alan and Sandra Gerry Metastasis and Tumor Ecosystems Center, Memorial Sloan Kettering Cancer Center
Ramon Parsons, M.D., Ph.D.
Director, Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai
Kornelia Polyak, M.D., Ph.D.
Professor of Medicine, Department of Medicine, Medical Oncology, Harvard Medical School and Dana-Farber Cancer Institute
Martine Roussel, Ph.D.
Professor, Department of Genetics and Tumor Cell Biology, St. Jude Children’s Research Hospital
Reuben Shaw, Ph.D.
Director, Salk Institute Cancer Center
Robert Winn, M.D.
Director and Lipman Chair in Oncology, VCU Massey Cancer Center
Past Directors

Dr. James Watson
1987 – 1988

Dr. Richard Roberts
1988 – 1992

Dr. Bruce Stillman
1992 – 2016
Zombie cells: The intersection of cancer and aging
July 1, 2025
These zombies have more to say than just “brains.” Let's see what they might tell us about the secrets of youth, rejuvenation, health, and disease.
A new “link” to triple-negative breast cancer
June 30, 2025
There’s no effective treatment for the deadly disease. A discovery by CSHL Professor David Spector and grad student Wenbo Xu could point to the first.
A recipe to reverse cancer’s sweet tooth
June 16, 2025
Researchers reveal the first 3D structures of the kinase FN3K in different states, which may inform future therapeutics for a variety of cancers.
At the Lab: Catch me if you cancer
June 3, 2025
You’ve seen her on FOX and the cover of Newsday. Now, hear about the knowledge gaps that make her discovery so crucial and the noble goals driving it.
At the Lab: UTIs in women’s health
May 13, 2025
Why do breast cancer researchers focused on pregnancy start looking into urinary tract infections? The backstory is as intriguing as the story itself.
At the Lab: The nuclear speckle option
May 6, 2025
Speckles sound innocent enough, like little spots. But don’t be fooled. These tiny structures could someday have big implications for cancer care.
From finance to restaurateur to men’s health advocate
April 24, 2025
At just 48, Tom Milana was diagnosed with prostate cancer. 10 years later he is making a big impact on prostate cancer research and men’s health.
It’s not you—it’s cancer
April 10, 2025
CSHL Associate Professor Tobias Janowitz and colleagues discover the brain-body connection underlying apathy in the late stages of cancer.
In pancreatic cancer, a race against time
April 2, 2025
By the time it’s diagnosed, it’s often too late. Now, Cold Spring Harbor Laboratory researchers have found a way to intercept the deadly disease.
Cocktails & Chromosomes: The hidden world inside our cells
March 20, 2025
What are nuclear speckles and how might these mysterious structures impact your DNA? Get an inside look from CSHL Assistant Professor Kate Alexander.
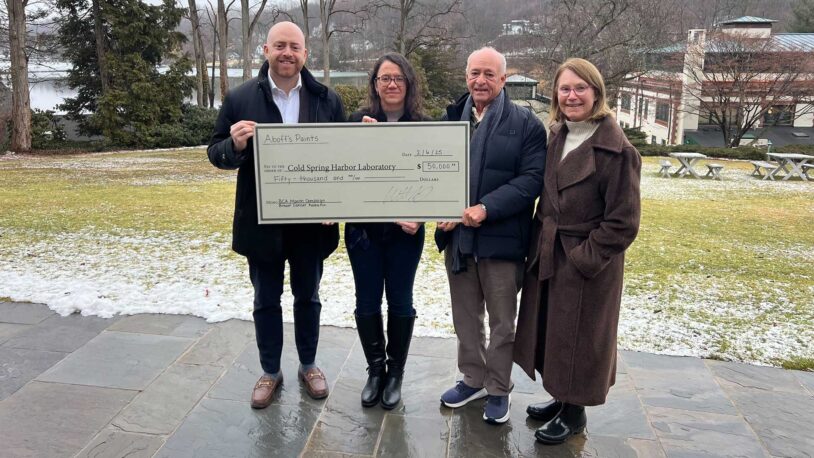
Aboff’s Paints donates $50K to CSHL breast cancer research
February 25, 2025
Assistant Professor Camila dos Santos and her colleagues in the CSHL Cancer Center thank the local family business for their continuing support.
CSHL receives $100K for leukemia research
February 24, 2025
The Don Monti Memorial Research Foundation’s donation supports the Zhang lab’s ongoing research on several variants of the deadly blood cancer.
Janowitz elected to ASCI
January 27, 2025
As a physician-scientist at CSHL, Tobias Janowitz has led early-phase clinical trials to develop therapeutic strategies for patients with cancer.
‘Is cancer immunotherapy right for me?’
January 21, 2025
It’s a question that only you and your doctor can answer. Research from CSHL’s Peter Westcott may explain why it does or doesn’t work in some cases.
A speckle of hope for cancer patients
January 2, 2025
Research from new CSHL Assistant Professor Katherine Alexander suggests that nuclear speckles could influence kidney cancer patient outcomes.
Cocktails & Chromosomes: Cancer’s identity crisis
December 19, 2024
CSHL Professor Christopher Vakoc shares his research with a standing-room-only crowd at Industry bar in Huntington, NY.
Women’s lives, breast health, and cancer
December 17, 2024
A panel discussion featuring CSHL Associate Professor Camila dos Santos and three clinical professionals specializing in women’s health.
The Microprocessor inside you
December 2, 2024
Stunning new images from CSHL’s Joshua-Tor lab reveal how the protein complex interacts with differently shaped primary microRNAs.
CSHL and Range ETFs launch new partnership
November 21, 2024
The affiliation was announced during a historic event at the Nasdaq stock market, highlighting the importance of cancer research.
Cocktails & Chromosomes: Long-lived creatures of the night
November 19, 2024
Bat genes may hold the keys to aging, immunity, and cancer. Find out how, as we relive a special Halloween night event with CSHL’s Dick McCombie.
Flipping cancer’s off switch
November 7, 2024
CSHL Professor Christopher Vakoc and postdoc Olaf Klingbeil have found a potential drug target for some of the most common and deadly cancers.
Adrian Krainer wins Albany Prize for biomedical research
October 8, 2024
A pioneer in the burgeoning field of RNA therapeutics, Krainer has now received America’s second-highest prize in medicine.
At the Lab Season 1 Research Rewind: Cancer
October 8, 2024
As the first season of our new podcast winds down, we’re revisiting all of our episodes with a focus on CSHL’s cutting-edge cancer research.
How does cancer spread? Follow the map
September 25, 2024
CSHL Professor Adam Siepel and postdoc Armin Scheben use genetic barcodes to map how prostate cancer spreads.
The curious immune cells caught between worlds
September 24, 2024
CSHL’s Hannah Meyer shows innate-like T cells mature differently in humans and mice. Her discovery could improve preclinical immunotherapy studies.
Cocktails & Chromosomes: How the brain and body communicate
September 19, 2024
Hungry? How do you know? To answer questions like this, CSHL’s Jeremy Borniger taps into the circuits controlling brain-body communication.
At the Lab Episode 24: Putting the brakes on brain cancer
September 17, 2024
CSHL Professor Alea Mills compares the deadly brain cancer glioblastoma to a car with its brakes cut. Her lab works to reattach them.
CSHL & Aboff’s Paints announce Paint & Donate campaign
September 9, 2024
Proceeds from every gallon of premium paint sold at Aboff’s Paints in October will go to support CSHL’s breast cancer research programs.
At the Lab Episode 22: Outmuscling cancer
September 3, 2024
After 10 years, CSHL has made a breakthrough in the study of RMS, a rare pediatric cancer. How we got here is a story of innovation and perseverance.
Humans of Banbury: Interview with Alessandra Luchini
August 30, 2024
An interview with June 2022's "Perinatal Transmission of Lyme Disease" meeting participant Alessandra Luchini, Ph.D.
How breast cancer goes hungry
August 28, 2024
CSHL Assistant Professor Michael Lukey has identified a source of breast cancer’s backup food supplies—and a way to block access.
Cocktails & Chromosomes: The whole-body health spectrum
August 22, 2024
How can a tiny tumor, often no larger than a penny, have such a big impact? CSHL’s Tobias Janowitz unpacks a complicated but highly relatable issue.
At the Lab Episode 20: Cleanup on IL-6
August 20, 2024
CSHL Professor Bo Li set out to study a tiny group of neurons involved in cancer cachexia. What he found astounded even him.
At the Lab Episode 18: The stress of cancer
August 6, 2024
A sobering conversation on a breakthrough discovery with potentially significant implications for cancer patients everywhere.
Katherine Alexander joins CSHL cancer faculty
August 5, 2024
Her team will explore mysterious cellular structures known as nuclear speckles and their role in diseases such as cancer.
A century of women’s health breakthroughs
July 22, 2024
Oscar Riddle identified the hormone behind lactation in 1933. The discovery at CSHL continues to inspire research on women’s health and breast cancer.
Cocktails & Chromosomes: Why do we age?
July 18, 2024
Humans have asked this question since the dawn of civilization. Today, we have the answer. Find out about this and more from CSHL’s Corina Amor Vegas.
One Experiment: Brain-body’s feathery display
July 15, 2024
An angry peacock is no joke. Like the colorful bird and its tall tail feathers, cancer biology can make for some eye-catching images.
Recapping CSHL’s 88th Symposium
July 10, 2024
Thought leaders from around the world came together at CSHL to share and discuss the latest research in brain-body physiology.
How to stop cancer cachexia? Start at the top
July 8, 2024
Professor Bo Li and a team of collaborators from four CSHL labs have discovered a new potential drug target for the lethal wasting disease.
CSHL’s 30th Annual Golf Tournament raises nearly $700,000
July 3, 2024
It was another record-breaker for the summer fundraiser, which this year honored Paul Paternoster, a longtime supporter of sarcoma research.
At the Lab Episode 13: A more sustainable chemistry
July 2, 2024
For this week’s podcast, CSHL Professor John Moses bridges the gap between chemistry and biology in less than three minutes.
At the Lab Episode 12: Cancer cops
June 25, 2024
In this week’s podcast, CSHL Assistant Professor Semir Beyaz reveals how metastatic breast cancer ‘corrupts’ the body’s immune system.
Pancreatic cancer’s cellular amnesia
June 17, 2024
New study from CSHL Professor Christopher Vakoc and former postdoc Diogo Maia-Silva shows how basal-like cancer cells lose their original identity.
Wrexham University awards John Moses Honorary Fellowship
June 13, 2024
CSHL Professor John Moses returned to his hometown of Wrexham, Wales, where he was recognized for his contributions to science.
RNA splicing’s spotters
June 10, 2024
RNA therapeutics pioneer CSHL Professor Adrian Krainer has discovered a link between two important regulator proteins, SRSF1 and DDX23.
At the Lab Episode 10: The time of our lives
June 4, 2024
“You wouldn’t start making the fingernails on an arm until you had started to make the arm,” says CSHL’s Christopher Hammell. How’s that for a visual?
The Louise Lindsay Read paperbark maple
May 30, 2024
Like the Chinese maple outside Blackford Hall, the roots of CSHL’s broad intellectual family tree run deep.
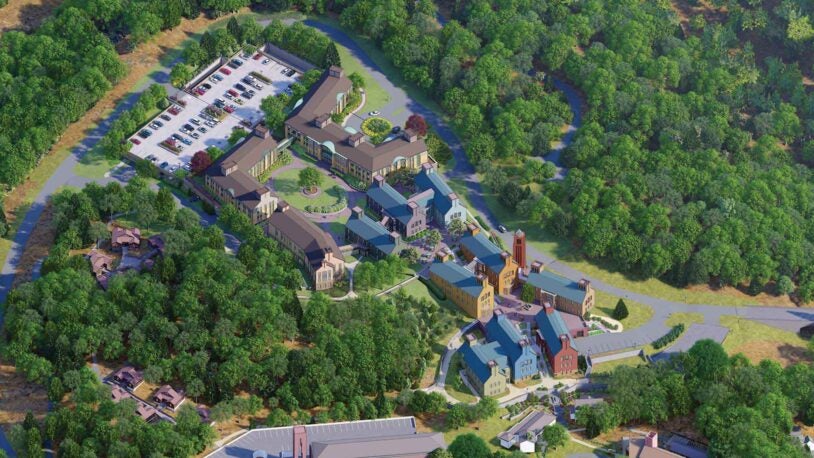
Tour CSHL’s Foundations for the Future project
May 9, 2024
Join New York Governor Kathy Hochul and CSHL President & CEO Bruce Stillman for an aerial tour of CSHL’s Foundations for the Future expansion project.
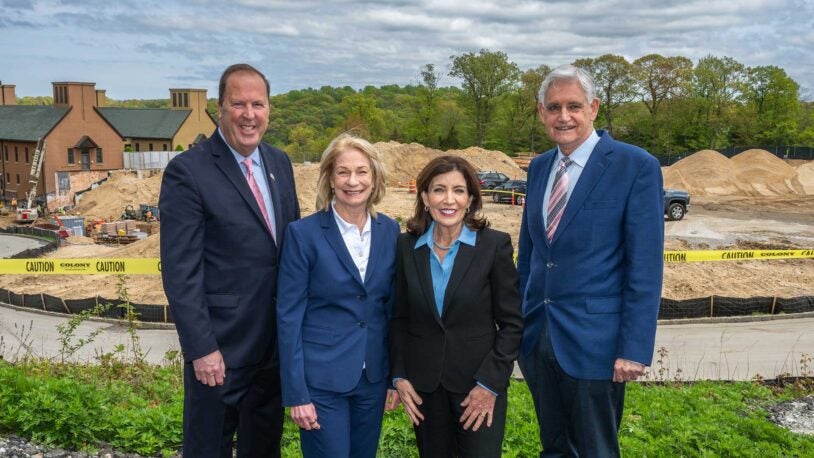
Governor Hochul announces $15 million for CSHL expansion
May 9, 2024
The funding will help pay for a new pancreatic cancer center—part of CSHL’s Foundations for the Future project.
A link between breast changes and . . . UTIs?
May 2, 2024
CSHL breast cancer researchers, led by Associate Professor Camila dos Santos, made a surprise discovery while investigating urinary tract infections.
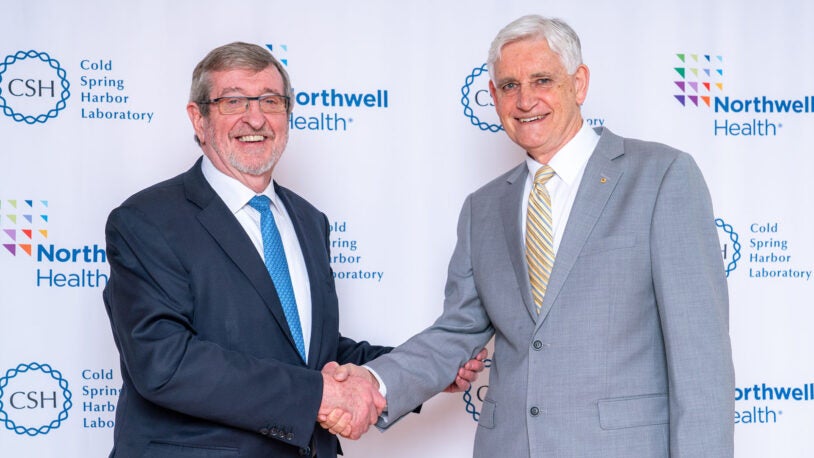
CSHL and Northwell Health extend strategic affiliation
April 29, 2024
Historic agreement aims to translate basic bioscience into a clinical setting, providing cancer patients greater access to personalized healthcare.
Tuveson elected to American Academy of Arts & Sciences
April 24, 2024
CSHL Cancer Center Director and Professor David Tuveson joins an elite membership roster whose historic ranks include names like Darwin and Einstein.
Nobel laureate honored at CSHL chemistry symposium
April 15, 2024
“The Future of Click Chemistry” brought together two-time Nobelist K. Barry Sharpless with his former apprentices John Moses and David Tuveson.
Click, click, boom—150 new molecules
April 4, 2024
CSHL Professor John Moses premieres an expansive line of new click chemistry products, uncovering leads for better antibiotics and cancer drugs.
One Experiment: Pancreatic mucus
March 26, 2024
That’s not the Starship Enterprise burning up in space. It’s an up-close look at precancerous pancreatic lesions and the mucus they produce.
Better cancer trials could be around the corner
March 21, 2024
CSHL researchers have developed a new data-driven approach that cancer centers may use to expand patient recruitment efforts.
Women’s health quiz
March 19, 2024
CSHL research has yielded insights into a number of women’s health topics, from menopause to breast cancer. Take this quiz to see how far we’ve come.
New hope in the fight against neurofibromatosis
March 18, 2024
A partnership between CSHL and the Penny’s Flight Foundation aims to find a cure for NF1, the world’s most common single-gene neurological disorder.
Cocktails & Chromosomes: Molecules to change the world
March 14, 2024
What is CSHL’s John Moses doing with that glowing liquid? Watch our expert chemist get a reaction from the crowd at Industry bar in Huntington, NY.
Women in science on women’s health
March 8, 2024
CSHL’s Camila dos Santos and Jessica Tollkuhn offer empowering insights into breast cancer prevention, pregnancy, menopause, and hormone therapy.
Pancreatic cancer lives on mucus
February 28, 2024
Mucus is not just snot. CSHL scientists have discovered some pancreatic cancer cells depend on it. The finding may open a new therapeutic avenue.
Can AI uncover breast cancer risk factors?
February 26, 2024
This question lies at the heart of a new interdisciplinary collaboration between CSHL’s Camila dos Santos and Peter Koo.
Chronic stress spreads cancer … here’s how
February 22, 2024
CSHL researchers have discovered a new link between chronic stress and cancer metastasis, providing a possible path forward for new treatments.
Pancreatic cancer hijacks a brain-building protein
February 14, 2024
CSHL researchers have found that EN-1, a protein involved in neurodevelopment, can help pancreatic cancer spread throughout the body.
From novel discovery to anti-aging therapy
February 8, 2024
Former CSHL Fellow Carol Greider’s Nobel-winning research has led to new cancer treatments. Now, it’s helping us unravel the mysteries of aging.
How diet may impact cancer and possible treatments
February 1, 2024
Researchers at the CSHL Cancer Center study the links between disease and nutrition in hopes of uncovering new treatment and prevention strategies.
The fountain of youth is … a T cell?
January 24, 2024
CSHL's Corina Amor Vegas discovers that CAR T cells can act as a “living” drug, causing young mice to age more slowly and old mice to rejuvenate.
From cancer genetics to cancer treatments
January 18, 2024
One cancer gene, one cancer genome, two Cold Spring Harbor Laboratory discoveries that helped shape the face of modern cancer medicine.
Joshua-Tor named CSHL Director of Research
January 2, 2024
The Cold Spring Harbor Laboratory professor and HHMI investigator steps into her new role effective January 2, 2024.
Cocktails & Chromosomes: You are what you eat
December 14, 2023
The question is how! Whet your appetite for discovery with this mouthwatering talk on diet and nutrition from CSHL’s Semir Beyaz.
Can we crack this cancer’s immune response?
November 29, 2023
CSHL research brings scientists closer to understanding how pancreatic cancer interacts with our immune system and why immunotherapy hasn’t worked.
CSHL celebrates 50th anniversary of McClintock Laboratory
November 20, 2023
Fifty years ago, CSHL honored Barbara McClintock by dedicating a building in her name. Today, it is home to four innovative cancer labs.
Foundations for the Future: Blueprint for tomorrow
November 13, 2023
Cold Spring Harbor Laboratory’s Advancement team provides an overview of the institution’s newly launched seven-acre expansion project.
First look: Whole-body gene expression
November 6, 2023
For the first time, scientists at CSHL have observed gene expression as it occurs throughout an animal. See life take shape in front of your eyes.
New antibody could target breast cancers
October 30, 2023
CSHL Professor Nicholas Tonks’ team has discovered a new way to target PTPRD, an enzyme that may help some breast cancers spread.
Vakoc receives Excellence in Healthcare Award
October 26, 2023
The award recognizes his breakthrough sarcoma research. This is the second year in a row that the LI Herald has honored a member of the CSHL faculty.
Test your breast cancer awareness
October 18, 2023
Awareness is key to prevention and potential future treatments. Take this quiz to find out about the latest in breast cancer research at CSHL.
Cocktails & Chromosomes: Breast cancer awareness edition
October 13, 2023
CSHL Associate Professor Camila dos Santos unpacks the science of breast cancer prevention at Industry bar in Huntington, NY.
CSHL announces AACR Cancer Centers Alliance
September 18, 2023
The new initiative, led by CSHL Professor David Tuveson, seeks to unite the nation’s NCI-designated cancer centers to help save more patients’ lives.
In cancer immunotherapy, more isn’t necessarily better
September 14, 2023
CSHL Assistant Professor Peter Westcott may have discovered why immunotherapy often fails in certain colon cancer treatments.
These worms have rhythm
September 5, 2023
Observing gene expression in real time, CSHL scientists identified four molecules the C. elegans worm relies on to set the tempo of its development.
Once rhabdomyosarcoma, now muscle
August 28, 2023
“Every successful medicine has its origin story,” says CSHL’s Christopher Vakoc. “And research like this is the soil from which new drugs are born.”
CSHL makes a splash with Swim Across America
August 21, 2023
Since 1987, the charity swim has raised over $100 million for cancer research. Here, CSHL Assistant Professor Semir Beyaz voices his support.
Growing human organs in the lab
August 3, 2023
It’s not something out of science-fiction. It’s a real biomedical breakthrough. This video, with organoid expert Dr. James Wells, shows how it works.
Metastatic breast cancer’s Trojan horse
July 26, 2023
CSHL and Massachusetts General Hospital researchers uncover how metastatic breast cancer slips past the immune system’s defenses in mice.
Lingbo Zhang wins National Institutes of Health MERIT Award
June 15, 2023
The highly prestigious award will support Zhang’s research on the role of nutrients and other environmental factors in blood cancer development.
Humans of Banbury: Interview with Marcus Goncalves
May 31, 2023
An interview with March 2023's "The Future of Investigational Medicine" meeting participant Marcus Goncalves, M.D., Ph.D.
Humans of Banbury: Interview with Naranjargal Dashdorj
May 12, 2023
An interview with March 2023's "The Future of Investigational Medicine" meeting participant Naranjargal Dashdorj, M.D., Ph.D.
Your diet is your future
May 3, 2023
In this video, CSHL Assistant Professor Semir Beyaz shows how what you eat continues to influence your health long after you’ve digested it.
Identifying cancer genes’ multiple personalities
April 10, 2023
The same genes can cause different subtypes of tumors. Now, CSHL can recreate them in the lab. The approach could lead to new cancer treatments.
Shrinking tumors with electricity
March 8, 2023
Step inside the lab of CSHL Associate Professor Jeremy C. Borniger, where he and his team are rewiring the nervous system to combat cancer cells.
The shocking new research making cancer nervous
March 8, 2023
Imagine an electronic device that can eliminate tumors. In our exclusive interview, Jeremy Borniger offers an inside look at this exciting new field.
CSHL to receive $2 million in federal budget for 2023
January 5, 2023
The money will help the Laboratory purchase new supercomputers and artificial intelligence equipment used in cancer research.
Unlocking cancer’s ancestry
December 27, 2022
New software may help reveal the complete connections between ancestry and cancer, which could lead to better, more personalized treatments.
David Tuveson wins 2022 Luminary Award
November 17, 2022
The award, presented on World Pancreatic Cancer Day, honors Tuveson for his efforts to find a cure and improve patients’ lives.
David Tuveson elected to National Academy of Medicine
October 17, 2022
Tuveson was recognized for his pioneering cancer organoid research that led to the development of the first mouse models for pancreatic cancer.
A universal cancer treatment?
October 13, 2022
A medicine that disrupts the DNA replication of cancer cells may be within reach.
Tree lighting for pediatric cancer awareness
September 30, 2022
CSHL researchers and local donors came together for the first annual tree lighting ceremony to commemorate Pediatric Cancer Research Awareness Month.
Immunologist Peter Westcott joins CSHL faculty
September 12, 2022
The Westcott lab explores how the immune system shapes tumor evolution, with the goal of developing new cancer immunotherapies.
Director, David Tuveson, M.D., Ph.D.
Deputy Director, Research, Chris Vakoc, Ph.D.
Cancer researchers at CSHL are using cutting-edge technology in innovative and collaborative studies to explore the basic biology underlying the disease. Our research can be divided into three main focus areas:
| Cancer Genetics and Genomics | |
| Cellular Communication in Cancer | |
| Gene Regulation and Inheritance |
The CSHL Cancer Center has long been a leader in basic research, exploring the fundamental pathways and molecules that enable life. Now, Cancer Center researchers are applying these groundbreaking discoveries to the development of new treatments and better diagnostics for cancer.
While maintaining its focus on exceptional basic science, the CSHL Cancer Center is also expanding translational research. The Lab has partnered with a leading healthcare system to increase preclinical research at the Lab and facilitate clinical trials based on basic science discoveries. At the same time, the next generation of doctors can experience basic research firsthand through translational training opportunities in the CSHL Cancer Center that are designed to bridge the gap between the lab and the clinic.

Katherine Alexander
Your genetic material, the DNA, is housed in a small compartment within your cells, called the cell nucleus. While famous as the cell’s DNA container, the nucleus is not just DNA. It also contains non-DNA substructures that intermingle with DNA. We study how DNA interfaces with these substructures and why this organization is important.

Corina Amor Vegas
As we age our body accumulates damaged “senescent” cells that our immune system is no longer able to effectively eliminate. Senescent cells are responsible for the development of aging and age-related diseases like cancer or fibrosis. My group studies how senescent cells evade the immune system thereby identifying new therapeutic approaches.

Semir Beyaz
Are you really what you eat? Our goal is to uncover the precise mechanisms that link nutrition to organismal health and disease states at the cellular and molecular level. A particular focus in our lab is to understand how dietary perturbations affect the immune system and contribute to the risk of diseases that are associated with immune dysfunction such as cancer.
Our laboratory deciphers how nutritional cues and metabolic–epigenetic cross-talk sculpt cellular networks that sustain healthy tissue function—and how their disruption drives maladaptive remodeling in diseases such as colon and endometrial cancers as well as in endometriosis. We integrate genomic, transcriptomic, and epigenomic profiling with metabolic measurements, biochemical assays, and microbiome analyses to deconstruct the signaling circuits underlying these outcomes. Anchoring our studies in patient-derived tissue specimens and organoid platforms ensures every mechanistic insight is rooted in human biology—and directly informs the discovery of biomarkers and therapeutic targets for prevention, early diagnosis, and treatment of these debilitating diseases.

Jeremy C. Borniger
Patients with cancer frequently experience debilitating symptoms that can impair quality of life and reduce odds of survival. These include drastic changes in appetite, sleep/wake cycles, cognitive function, and pain, among others. Our lab aims to uncover mechanistic interactions between the brain and cancer that drive these phenomena. Reciprocally, we investigate how manipulation of specific brain circuits influences cancer processes in the body.

Jeff Boyd
My research interests are in the molecular genetics, genetics, and genomics of gynecologic and breast cancers. Currently I am focused on the early natural histories of ovarian carcinoma and metastatic breast cancer, the genomics of ovarian cancer stem/progenitor cells, and the hypothesis that most breast cancers result from polygenic susceptibility.
Nyasha Chambwe
My research focuses on identifying the genetic and molecular features of cancers that differ across racial and ethnic groups, and the extent to which these differences reveal or explain race and ethnicity-based cancer health disparities.

Kenneth Chang
RNA interference (RNAi) and CRISPR are widely used to functionally investigate mammalian genomes. It is our goal to develop and optimize these gene perturbation platforms to improve their effectiveness in understanding the biology of diseases.

Lucas Cheadle
The trillions of connections between brain cells enable complex thought and behavior. These connections are wired with great precision through both genetics and in response to an organism’s experiences. Our lab seeks to understand how experiences engage specialized immune cells called microglia to shape the connectivity and function of the brain. We are further interested in how impairments in these processes can contribute to neurodevelopmental disorders such as autism.

Paolo Cifani
We develop innovative mass spectrometry-based approaches to measure how protein activities are regulated under physiologic conditions and in pathological states.

Alexander Dobin
Next generation sequencing technologies revolutionized many areas of genetics and molecular biology, enabling quantitative analyses of the entire genomes and paving the way for Personalized Medicine. We develop novel statistical methods and computational algorithms for multi-omics processing and integration, and leverage Big Genomic Data to elucidate various problems in precision health, such as genetic and epigenetic mechanisms of cancer development and progression, and clinical impact of functional variants.

Camila dos Santos
Among the changes that occur during pregnancy, those affecting the breasts have been found to subsequently modify breast cancer risk. My laboratory investigates how the signals present during pregnancy permanently alter the way gene expression is controlled and how these changes affect normal and malignant mammary development.

Douglas Fearon
I’m studying how to harness the power of the immune system to fight cancer. Our underlying premise is that the microenvironment within a tumor suppresses the immune system. We have found a way to eliminate this suppression in the mouse model of pancreatic cancer, which has led to development of a drug for human pancreatic cancer that will enter phase 1 clinical trials in 2015.

Hiro Furukawa
The nervous system transmits information by passing chemical signals from one nerve cell to the others. This signal transmission relies on a variety of proteins to receive and transmit the chemical signals. My group studies the structure and function of neurotransmitter receptors and ion channels that regulate fundamental neuronal activities.

Sepideh Gholami
Dr. Sepideh Gholami M.D., M.A.S. is a board-certified surgeon scientist with dual fellowship training in Complex General Surgical Oncology and Hepatopancreatobiliary Surgery. Dr. Gholami serves as the Director of the Liver Multidisciplinary Clinic, Hepatic Artery Infusion Pump Program, and Translational Research in Surgical Oncology at Northwell Health. She has focused her efforts on building a multidisciplinary liver surgery program with liver-directed therapies/regional therapies, including a hepatic artery infusion pump program for patients with hepatobiliary and metastatic malignancies. Dr. Gholami’s mission is to diversify and improve the research and clinical trial portfolio at Northwell Health Cancer Institute. She also has a joint appointment as an Adjunct Associate Professor at Cold Spring Harbor Laboratory.

Thomas Gingeras
Only a small portion of the RNAs encoded in any genome are used to make proteins. My lab investigates what these noncoding RNAs (ncRNAs) do within and outside of cells, where regulators of their expression are located in the genome. This is particularly important in cancer. Our laboratory works on endometrial cancer and its relationship to age and obesity.

Sara Goodwin
I work on adapting and developing new methods/techniques for genome and transcriptome sequencing.

Christopher Hammell
As organisms develop, genes turn on and off with a precise order and timing, much like the order and duration of notes in a song. My group uses model organisms to understand the molecules that control the tempo of development. We also study how changes in the timing of gene expression contribute to diseases like cancer.

Tobias Janowitz
Cancer is a systemic disease. Using both laboratory and clinical research, my group investigates the connections between metabolism, endocrinology, and immunology to discover how the body’s response to a tumor can be used to improve treatment for patients with cancer.

Leemor Joshua-Tor
Our cells depend on thousands of proteins and nucleic acids that function as tiny machines: molecules that build, fold, cut, destroy, and transport all of the molecules essential for life. My group is discovering how these molecular machines work, looking at interactions between individual atoms to understand how they activate gene expression, DNA replication, and small RNA biology.

Justin Kinney
Research in the Kinney Lab combines mathematical theory, machine learning, and experiments in an effort to illuminate how cells control their genes. These efforts are advancing the fundamental understanding of biology and biophysics, as well as accelerating the discovery of new treatments for cancer and other diseases.

Peter Koo
Deep learning has the potential to make a significant impact in basic biology and cancer, but a major challenge is understanding the reasons behind their predictions. My research develops methods to interpret this powerful class of black box models, with a goal of elucidating data-driven insights into the underlying mechanisms of sequence-function relationships.

Adrian R. Krainer
Our DNA carries the instructions to manufacture all the proteins needed by a cell. After each gene is copied from DNA into RNA, the RNA message is "spliced" - an editing process involving precise cutting and pasting. I am interested in how splicing normally works, how it is altered in genetic diseases and cancer, and how we can correct these defects for therapy.

Alexander Krasnitz
Many types of cancer display bewildering intra-tumor heterogeneity on a cellular and molecular level, with aggressive malignant cell populations found alongside normal tissue and infiltrating immune cells. I am developing mathematical and statistical tools to disentangle tumor cell population structure, enabling an earlier and more accurate diagnosis of the disease and better-informed clinical decisions.

Dan Levy
We have recently come to appreciate that many unrelated diseases, such as autism, congenital heart disease and cancer, are derived from rare and unique mutations, many of which are not inherited but instead occur spontaneously. I am generating algorithms to analyze massive datasets comprising thousands of affected families to identify disease-causing mutations.

Michael Lukey
Tumor growth depends upon cancer cells acquiring nutrients from their environment and using these molecules to fuel proliferation. My group studies the nature and regulation of metabolic adaptation during tumorigenesis and metastasis, with the intention of identifying metabolic vulnerabilities that can be targeted for cancer therapy.

Scott Lyons
I provide collaborative research support to CSHL researchers in the area of preclinical in vivo imaging. This includes access to a comprehensive range of imaging modalities, as well as provision of experimental guidance, training and imaging reagents. In addition, my lab develops new and impactful ways to image aspects of in vivo tumor biology that are broadly relevant to the development of new therapeutics and the research interests of the CSHL Cancer Center.

Rob Martienssen
Chromosomes are covered with chemical modifications that help control gene expression. I study this secondary genetic code - the epigenome - and how it is guided by small mobile RNAs in plants and fission yeast. Our discoveries impact plant breeding and human health, and we use this and other genomic information to improve aquatic plants as a source of bioenergy.

David McCandlish
Some mutations are harmful but others are benign. How can we predict the effects of mutations, both singly and in combination? Using data from experiments that simultaneously measure the effects of thousands of mutations, I develop computational tools to predict the functional impact of mutations and apply these tools to problems in protein design, molecular evolution, and cancer.

W. Richard McCombie
Over the last two decades, revolutionary improvements in DNA sequencing technology have made it faster, more accurate, and much cheaper. We are now able to sequence up to 10 trillion DNA letters in just one month. I harness these technological advancements to assemble genomes for a variety of organisms and probe the genetic basis of neurological disorders, including autism and schizophrenia, better understand cancer progression and understand the complex structures of the genomes of higher plants.

Hannah Meyer
A properly functioning immune system must be able to recognize diseased cells and foreign invaders among the multitude of healthy cells in the body. This ability is essential to both prevent autoimmune diseases and fight infections and cancer. We study how a specific type of immune cells, known as T cells, are educated to make this distinction during development.

Alea A. Mills
Cells employ stringent controls to ensure that genes are turned on and off at the correct time and place. Accurate gene expression relies on several levels of regulation, including how DNA and its associated molecules are packed together. I study the diseases arising from defects in these control systems, such as aging and cancer.

Lopa Mishra
My research focuses on the continuum of science-driven clinical care by working on novel therapies and improved clinical outcomes, honing liver disease, metabolism/alcohol, obesity/addiction gastrointestinal cancers, inflammatory bowel disease, and neural regulation of disease and cancer, which links to the field of bioelectronic medicine.

John Moses
My group uses click chemistry to study biological systems at the molecular level. We develop and exploit powerful bond-forming click reactions that enable the rapid synthesis of small functional molecules, including cancer drugs and chemical probes. We apply these novel molecular tools in multidisciplinary discovery projects spanning the fields of biology and chemistry.

Jon Preall
Developing single-cell genomics technologies for applications related to cancer progression, immune surveillance, and discovery of rare novel cell types and transcriptional programs.

Andrea Schorn
Transposable elements make up half of our DNA. They control gene expression and have been a major evolutionary force in all organisms. The Schorn lab investigates how small RNAs identify and silence transposable elements when they become active during development and disease.

Adam Siepel
I am a computer scientist who is fascinated by the challenge of making sense of vast quantities of genetic data. My research group focuses in particular on questions involving molecular evolution and transcriptional regulation, with applications to cancer and other diseases as well as to plant breeding and agriculture.

David L. Spector
The immense amount of DNA, RNA and proteins that contribute to our genetic programs are precisely organized inside the cell's nucleus. My group studies how nuclear organization impacts gene regulation, and how misregulation of non-coding RNAs contributes to human diseases such as cancer.

Bruce Stillman
Every time a cell divides, it must accurately copy its DNA. With 3 billion “letters” in the human genome, this is no small task. My studies reveal the many steps and molecular actors involved, as well as how errors in DNA replication are involved in diseases that range from cancer to rare genetic disorders.

Jessica Tollkuhn
My lab studies how estrogen and testosterone regulate gene expression in the brain. The receptors for these steroid hormones directly bind DNA to turn genes on or off. We have found that sex differences in gene expression are a dynamic readout of hormone actions across the lifespan. We aim to understand how these hormone-regulated genes contribute to sex-variable biology, behavior, and disease risk.

Nicholas Tonks
Cells must constantly react to what is happening around them, adapting to changes in neighboring cells or the environment. I study the signals that cells use to exchange information with their surroundings. Our group is finding drugs that target these signals and thus can treat diabetes, obesity, cancer, and autism spectrum disorders.

Kevin Tracey
The major focus of my research is the molecular basis of inflammation and identifying the mechanisms by which neurons control the immune system.

Lloyd Trotman
We pioneered generation of a unique genetic mouse model for therapy and analysis of metastatic prostate cancer. Recently, we developed 3-dimensional whole organ imaging technology that allows us to visualize cancer and metastatic progression in its native environment and at single cell resolution. Now, we use this platform to understand the role of nerves in tumor metastasis, and to develop novel therapeutic interventions against lethal disease.

David Tuveson
Pancreatic cancer is an extremely lethal malignancy. On average, patients who are diagnosed with pancreatic cancer succumb to the disease within 6 months. Research is the only way to defeat pancreatic cancer. My lab is making progress toward finding a cure by detecting the disease earlier and designing novel therapeutic approaches.

Chris Vakoc
Cancer cells achieve their pathogenicity by changing which genes are on and off. To maintain these changes in gene expression, cancer cells rely on proteins that interact with DNA or modify chromatin. My group investigates how such factors sustain the aberrant capabilities of cancer cells, thereby identifying new therapeutic targets.

Linda Van Aelst
Normal cell function relies on coordinated communication between all the different parts of the cell. These communication signals control what a cell does, what shape it takes, and how it interacts with other cells. I study these signaling networks to understand how they guard against cancer and neurological disorders.

Erika Tse-Luen Wee
Develop and implement state-of-the-art fluorescence imaging and analysis techniques to quantify cell and tissue samples' structure and function.

Peter Westcott
The mutational processes that drive cancer also expose it to the immune system. Therapies that invigorate anticancer immunity can be astonishingly effective, but only in a subset of patients. We are developing powerful new strategies to study how the immune system and cancer coevolve, with the goal of expanding the curative potential of immunotherapy to more patients.

Michael Wigler
Devastating diseases like cancer and autism can be caused by spontaneous changes to our DNA—mutations first appearing in the child, or in our tissues as we age. We are developing methods to discover these changes in individuals, tumors, and even single cells, to promote early detection and treatments

Johannes Yeh
Cells orchestrate proteins to conduct cell-cell communications and environment sensing in order to execute physiological functions. My lab investigates the mechanisms by which dysregulated signals cause diseases such as cancer, and we are developing therapeutics based on these mechanisms.

Lingbo Zhang
The research in the Zhang laboratory centers on normal and malignant stem and progenitor cells in the hematopoietic system and decodes the role of metabolites in the tumor microenvironment, including nutrients and neurotransmitters, and their genetic effectors in regulating hematologic malignancies. The ultimate goal is to understand how environmental signals such as dietary and neuronal activities regulate stem and progenitor cell development and cancers.

Cold Spring Harbor Laboratory is an NCI-designated Cancer Center. As a basic research institution, CSHL does not treat patients. Information about individual cancers is available at the NCI CancerNet. Questions about CSHL’s cancer research program should be directed to our Communications Department.