Andrea Schorn
Assistant Professor
Cancer Center Member
Ph.D., Max-Delbrück Center for Molecular Medicine, Freie Universität Berlin, 2009
aschorn@cshl.edu | 516-367-8836
Faculty ProfileTransposable elements make up half of our DNA. They control gene expression and have been a major evolutionary force in all organisms. The Schorn lab investigates how small RNAs identify and silence transposable elements when they become active during development and disease.
‘Mobile genes’ or transposable elements, which are closely related to viruses, promote active gene expression in a selfish manner. These elements are usually buried in inactive, condensed DNA by their host to prevent mutagenic damage. However, both stem cells in the embryo and cancer cells undergo genome-wide reprogramming that re-activates silent transposable elements. My lab is exploring how the host recognizes transposons amongst thousands of genes and non-coding DNA and specifically restricts transposon mobility.
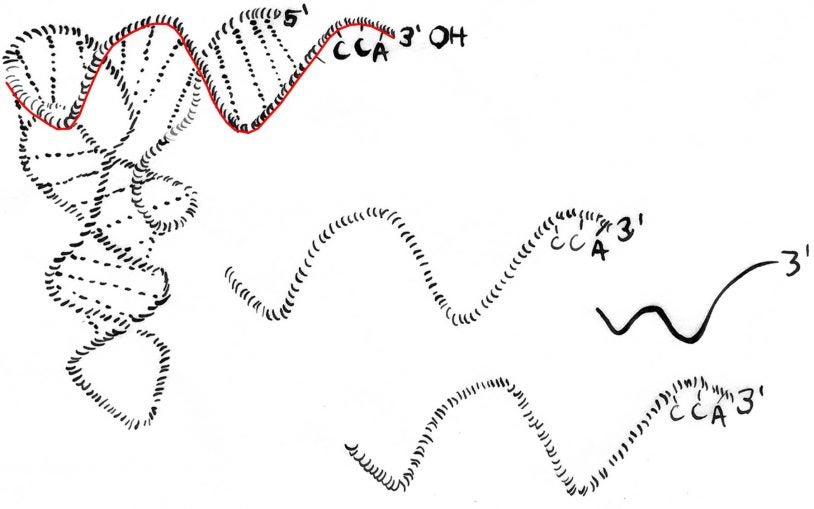
We found that a highly conserved 18 nucleotide sequence motif is the Achille’s heel of a wide-spread class of transposable elements that are closely related to retroviruses such as HIV. These retroelements initiate replication at the 18 nucleotide binding site using transfer RNA (tRNA), an essential RNA component of the cell. In turn, cells produce short fragments of tRNAs that we discovered inhibit this class of retroelements. These tRNA fragments are processed from mature tRNAs under yet unknown conditions and potentially protect many cell types in eukaryotes. We are investigating under which conditions cells produce this class of small RNAs and assessing their impact on development and pluripotency. tRNA fragments are an ancient link between the ‘RNA interference’ silencing machinery, transposons and genome stability, with potential roles in trans-generational inheritance and cancer.
The ancient viruses in human DNA
August 24, 2023
CSHL Assistant Professor Andrea Schorn takes us into her lab for a behind-the-scenes look at the parts of our genome that aren’t quite human.
Where did our DNA come from?
August 22, 2023
Half the human genome isn’t quite human. CSHL’s Andrea Schorn gives us the inside scoop on how our DNA turned ancient viruses into essential allies.
Andrea Schorn zooms in on small RNAs in the cell
April 29, 2019
Dr. Andrea Schorn joins CSHL faculty as research assistant professor. Her focus is exploring transposable elements in the genome of mammalian cells.
Newly identified small RNA fragments defend the genome when it’s ‘naked’
June 29, 2017
Fragments snipped from tRNAs protect embryonic stem cells while they’re being epigenetically reprogrammed.
LTR-Retrotransposon Control by tRNA-Derived Small RNAs
F1000Prime articles list
All Publications
Pseudouridine guides germline small RNA transport and epigenetic inheritance
6 Sep 2024 | Nature Structural & Molecular Biology
Herridge, Rowan; Dolata, Jakub; Migliori, Valentina; de Santis Alves, Cristiane; Borges, Filipe; Schorn, Andrea; van Ex, Frédéric; Lin, Ann; Bajczyk, Mateusz; Parent, Jean-Sebastien; Leonardi, Tommaso; Hendrick, Alan; Kouzarides, Tony; Martienssen, Robert;
Pseudouridine guides germline small RNA transport and epigenetic inheritance
3 Aug 2023 | bioRxiv
Herridge, Rowan; Dolata, Jakub; Migliori, Valentina; de Santis Alves, Cristiane; Borges, Filipe; Van Ex, Frédéric; Lin, Ann; Bajczyk, Mateusz; Leonardi, Tommaso; Hendrick, Alan; Schorn, Andrea; Kouzarides, Tony; Martienssen, Robert;
HMGXB4 Targets Sleeping Beauty Transposition to Germinal Stem Cells
14 Apr 2023 | International Journal of Molecular Sciences | 24(8):7283
Devaraj, Anantharam; Singh, Manvendra; Narayanavari, Suneel; Yong, Guo; Chen, Jiaxuan; Wang, Jichang; Becker, Mareike; Walisko, Oliver; Schorn, Andrea; Cseresznyés, Zoltán; Raskó, Tamás; Radscheit, Kathrin; Selbach, Matthias; Ivics, Zoltán; Izsvák, Zsuzsanna;
Dicer promotes genome stability via the bromodomain transcriptional co-activator BRD4
22 Feb 2022 | Nature Communications | 13(1):1001
Gutbrod, M; Roche, B; Steinberg, J; Lakhani, A; Chang, K; Schorn, A; Martienssen, R;
TET2 chemically modifies tRNAs and regulates tRNA fragment levels.
Jan 2021 | Nature Structural & Molecular Biology | 28(1):62-70
He, Chongsheng; Bozler, Julianna; Janssen, Kevin; Wilusz, Jeremy; Garcia, Benjamin; Schorn, Andrea; Bonasio, Roberto;
Endogenous Retroviruses Walk a Fine Line between Priming and Silencing.
23 Jul 2020 | Viruses | 12(8)
Cullen, Harrison; Schorn, Andrea;
Arabidopsis retrotransposon virus-like particles and their regulation by epigenetically activated small RNA.
Apr 2020 | Genome Research | 30(4):576-588
Lee, Seung; Ernst, Evan; Berube, Benjamin; Borges, Filipe; Parent, Jean-Sebastien; Ledon, Paul; Schorn, Andrea; Martienssen, Robert;
Getting in LINE with Replication
2 May 2019 | Molecular Cell | 74(3):415-417
Schorn, A; Martienssen, R;
Tie-Break: Host and Retrotransposons Play tRNA
Oct 2018 | Trends in Cell Biology | 28(10):793-806
Schorn, A; Martienssen, R;
LTR-Retrotransposon Control by tRNA-Derived Small RNAs
29 Jun 2017 | Cell | 170(1):61-71
Schorn, A; Gutbrod, M; LeBlanc, C; Martienssen, R;