Rob Martienssen
Professor & HHMI Investigator
William J. Matheson Professor
Cancer Center Member
Ph.D., Cambridge University, 1986
martiens@cshl.edu | 516-367-8322
Chromosomes are covered with chemical modifications that help control gene expression. I study this secondary genetic code - the epigenome - and how it is guided by small mobile RNAs in plants and fission yeast. Our discoveries impact plant breeding and human health, and we use this and other genomic information to improve aquatic plants as a source of bioenergy.
Epigenetic mechanisms of gene regulation—chemical and conformational changes to DNA and the chromatin that bundles it—have had an important impact on genome organization and inheritance and on cell fate. These mechanisms are conserved in eukaryotes and provide an additional layer of information superimposed on the genetic code. Robert Martienssen, a pioneer in the study of epigenetics, investigates mechanisms involved in gene regulation and stem cell fate in yeast and model plants including Arabidopsis and maize. He and his colleagues have shed light on a phenomenon called position-effect variegation, caused by inactivation of a gene positioned near densely packed chromosomal material called heterochromatin. They have discovered that small RNA molecules arising from repeating genetic sequences program that heterochromatin. Martienssen and colleagues have described a remarkable process by which “companion cells” to sperm in plant pollen grains provide them with instructions that protect sperm DNA from transposon damage. They found that some of these instructions, or epigenetic marks, could be inherited in the next generation. These marks, and the small RNA responsible that guide them, can sense the number of chromosomes inherited from pollen and may allow Arabidopsis, a flowering plant, to produce egg cells without meiosis, an important step toward a long-time goal of plant breeding: generating clonal offspring to perpetuate hybrid vigor. The lab has also shown that when RNA polymerase II has transcribed a stretch of DNA, the RNA interference mechanism causes the enzyme to release its hold on the DNA and fall away. This allows the replication fork to progress smoothly and the DNA strands to be copied; histone-modifying proteins, which follow right along, establish heterochromatin. Martienssen’s group also continues to work on problems related to the creation of plant-based biofuels. As part of a collaborative project to generate a high-quality full genome map of the oil palm plant, Martienssen and his colleagues identified a transposon whose modification controls the yield of oil palm trees. This discovery will increase yields and should lessen the environmental burden of oil palm production, which often threatens already endangered rainforest lands.
Martienssen named 2020 Royal Society winner
Rob Martienssen wins Martin Gibbs Medal for plant research
CSHL’s Rob Martienssen honored with prestigious Barbara McClintock Prize
Plant scientist Rob Martienssen receives prestigious appointment as HHMI-GBMF Investigator
“Breakthrough of the Year” for 2002
The food and fuel that farms itself
April 1, 2025
Duckweed can grow practically anywhere there’s sun and standing water. CSHL plant biologists identify the genes behind some of its most useful traits.
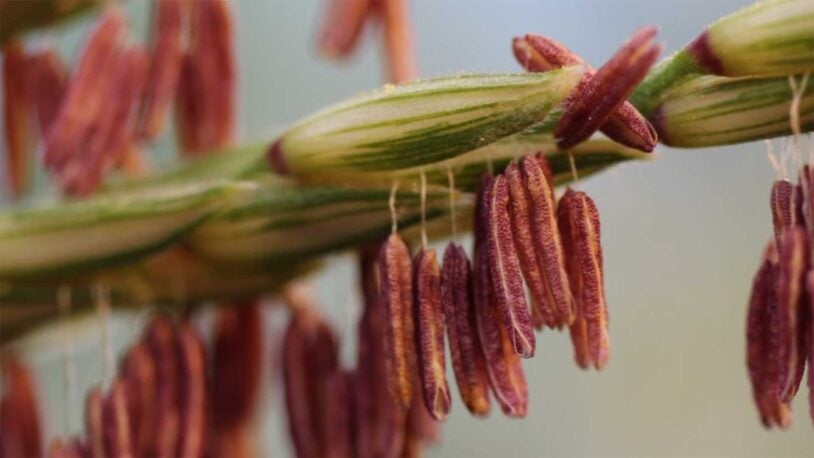
Corn’s ancient ancestors are calling
February 6, 2025
Cold Spring Harbor Laboratory Professors Thomas Gingeras and Rob Martienssen have launched a new genomic encyclopedia called MaizeCODE.
Big clues emerge in small RNA mystery
December 16, 2024
How does the body separate “self” from “nonself?” CSHL Professor Rob Martienssen uncovers fascinating new leads in plant pollen and mouse sperm.
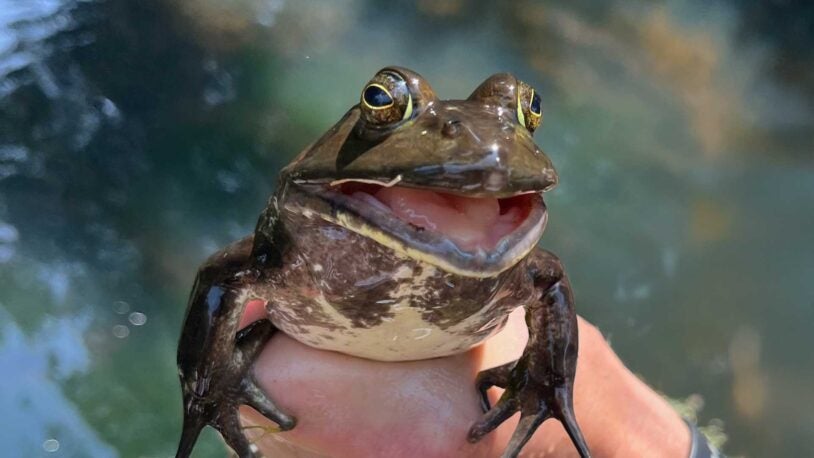
Frog Pond
November 27, 2024
These hopping amphibians aren’t the only animals that call the pond home. It’s a living snapshot of our area’s diverse ecosystem.
At the Lab Season 1 Research Rewind: Genetics
October 22, 2024
It’s the code for all life on Earth. This week At the Lab, we’re hacking it with the help of Cold Spring Harbor Laboratory’s geneticists.
Plants have a backup plan
October 3, 2024
CSHL’s Rob Martienssen and his team discovered how plants like Arabidopsis continue to reproduce even when things go wrong in chromosome division.
At the Lab Episode 25: How maize became corn
September 24, 2024
Cold Spring Harbor Laboratory solves a plant biology mystery some 4,000 years in the making. The implications may go far beyond vegetables.
Corn’s ‘missing link’
August 7, 2024
CSHL researchers have discovered a biological mechanism that may explain how corn spread so rapidly across the Americas 4,000 years ago.
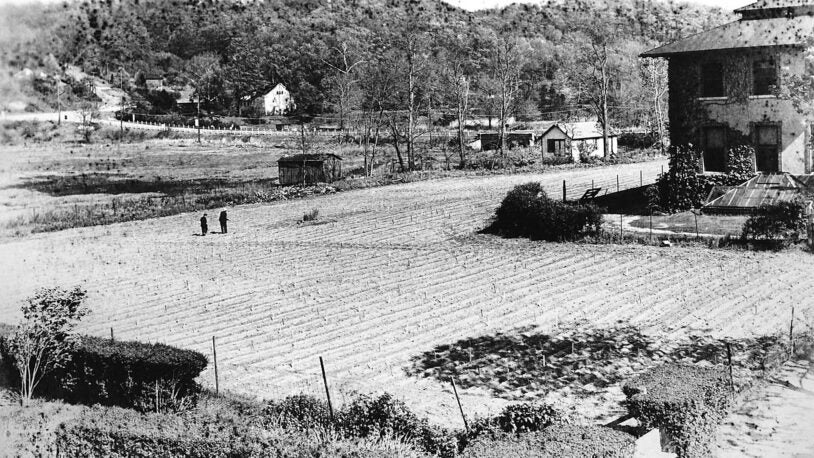
Barbara McClintock’s corn
June 16, 2024
You’ve heard about Barbara McClintock’s Nobel Prize-winning research on corn genetics, but what about the corn itself?
The CSHL School of Biological Sciences’ class of 2024
May 5, 2024
The School of Biological Sciences awarded Ph.D. degrees to 11 students this year. Here are some stories and reflections from their time at CSHL.
All Publications
Duckweed genomes and epigenomes underlie triploid hybridization and clonal reproduction
28 Mar 2025 | Current Biology
Ernst, Evan; Abramson, Bradley; Acosta, Kenneth; Hoang, Phuong; Mateo-Elizalde, Cristian; Schubert, Veit; Pasaribu, Buntora; Albert, Patrice; Hartwick, Nolan; Colt, Kelly; Aylward, Anthony; Ramu, Umamaheswari; Birchler, James; Schubert, Ingo; Lam, Eric; Michael, Todd; Martienssen, Robert;
MaizeCODE reveals bi-directionally expressed enhancers that harbor molecular signatures of maize domestication
30 Dec 2024 | Nature Communications | 15(1):10854
Cahn, Jonathan; Regulski, Michael; Lynn, Jason; Ernst, Evan; de Santis Alves, Cristiane; Ramakrishnan, Srividya; Chougule, Kapeel; Wei, Sharon; Lu, Zhenyuan; Xu, Xiaosa; Ramu, Umamaheswari; Drenkow, Jorg; Kramer, Melissa; Seetharam, Arun; Hufford, Matthew; McCombie, W; Ware, Doreen; Jackson, David; Schatz, Michael; Gingeras, Thomas; Martienssen, Robert;
Clr4SUV39H1 ubiquitination and non-coding RNA mediate transcriptional silencing of heterochromatin via Swi6 phase separation
30 Oct 2024 | Nature Communications | 15(1):9384
Kim, Hyun-Soo; Roche, Benjamin; Bhattacharjee, Sonali; Todeschini, Leila; Chang, An-Yun; Hammell, Christopher; Verdel, André; Martienssen, Robert;
Pseudouridine guides germline small RNA transport and epigenetic inheritance
6 Sep 2024 | Nature Structural & Molecular Biology
Herridge, Rowan; Dolata, Jakub; Migliori, Valentina; de Santis Alves, Cristiane; Borges, Filipe; Schorn, Andrea; van Ex, Frédéric; Lin, Ann; Bajczyk, Mateusz; Parent, Jean-Sebastien; Leonardi, Tommaso; Hendrick, Alan; Kouzarides, Tony; Martienssen, Robert;
Retrotransposon addiction promotes centromere function via epigenetically activated small RNAs
2 Sep 2024 | Nature Plants
Shimada, Atsushi; Cahn, Jonathan; Ernst, Evan; Lynn, Jason; Grimanelli, Daniel; Henderson, Ian; Kakutani, Tetsuji; Martienssen, Robert;
Leishmania major telomerase RNA knockout: From altered cell proliferation to decreased parasite infectivity
30 Aug 2024 | International Journal of Biological Macromolecules | :135150
de Oliveira, Beatriz; Shiburah, Mark; Assis, Luiz; Fontes, Veronica; Bisetegn, Habtye; de Oliveira Passos, Arthur; de Oliveira, Leilane; de Santis Alves, Cristiane; Ernst, Evan; Martienssen, Rob; Gallo-Francisco, Pedro; Giorgio, Selma; Batista, Marcos; de Nazaré Correia Soeiro, Maria; Menna-Barreto, Rubem; Aoki, Juliana; Coelho, Adriano; Cano, Maria;
Teosinte Pollen Drive guides maize diversification and domestication by RNAi
7 Aug 2024 | Nature
Berube, Benjamin; Ernst, Evan; Cahn, Jonathan; Roche, Benjamin; de Santis Alves, Cristiane; Lynn, Jason; Scheben, Armin; Grimanelli, Daniel; Siepel, Adam; Ross-Ibarra, Jeffrey; Kermicle, Jerry; Martienssen, Robert;
MaizeCODE reveals bi-directionally expressed enhancers that harbor molecular signatures of maize domestication
23 Feb 2024 | bioRxiv
Cahn, Jonathan; Regulski, Michael; Lynn, Jason; Ernst, Evan; Alves, Cristiane; Ramakrishnan, Srividya; Chougule, Kapeel; Wei, Sharon; Lu, Zhenyuan; Xu, Xiaosa; Drenkow, Jorg; Kramer, Melissa; Seetharam, Arun; Hufford, Matthew; McCombie, Richard; Ware, Doreen; Jackson, David; Schatz, Michael; Gingeras, Thomas; Martienssen, Robert;
Mechanisms of metabolic adaptation in the duckweed Lemna gibba: an integrated metabolic, transcriptomic and flux analysis
3 Oct 2023 | BMC Plant Biology | 23(1):458
Shi, Hai; Ernst, Evan; Heinzel, Nicolas; McCorkle, Sean; Rolletschek, Hardy; Borisjuk, Ljudmilla; Ortleb, Stefan; Martienssen, Robert; Shanklin, John; Schwender, Jorg;
Oncogene-like addiction to aneuploidy in human cancers
25 Aug 2023 | Science | 381(6660):eadg4521
Girish, Vishruth; Lakhani, Asad; Thompson, Sarah; Scaduto, Christine; Brown, Leanne; Hagenson, Ryan; Sausville, Erin; Mendelson, Brianna; Kandikuppa, Pranav; Lukow, Devon; Yuan, Monet; Stevens, Eric; Lee, Sophia; Schukken, Klaske; Akalu, Saron; Vasudevan, Anand; Zou, Charles; Salovska, Barbora; Li, Wenxue; Smith, Joan; Taylor, Alison; Martienssen, Robert; Liu, Yansheng; Sun, Ruping; Sheltzer, Jason;