Christopher Hammell
Professor
Cancer Center Member
Ph.D., Dartmouth Medical School, 2002
chammell@cshl.edu | 516-367-5207
Faculty ProfileAs organisms develop, genes turn on and off with a precise order and timing, much like the order and duration of notes in a song. My group uses model organisms to understand the molecules that control the tempo of development. We also study how changes in the timing of gene expression contribute to diseases like cancer.
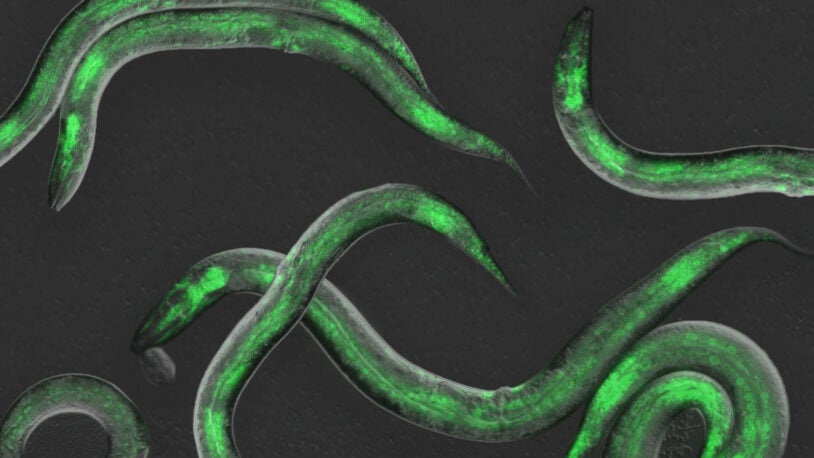
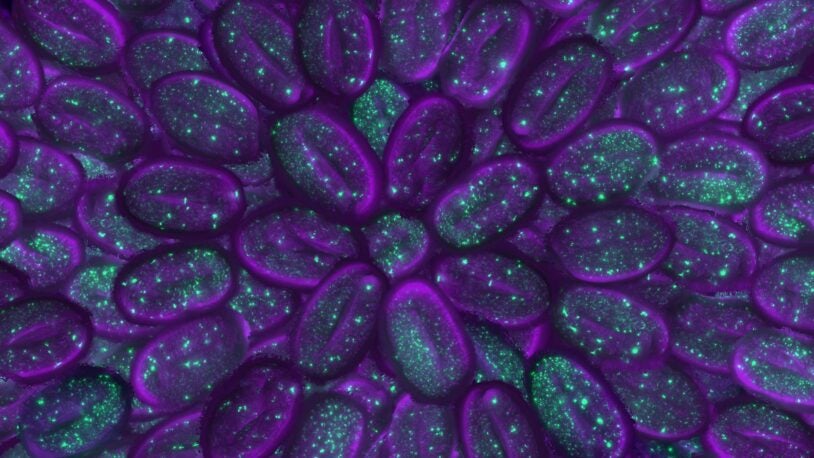
Christopher Hammell’s lab is interested in understanding gene regulatory processes that give rise to robust phenotypes associated with normal development in animals (specifically, how the timing of developmental processes is controlled) as well as the alterations in these pathways that give rise to diseases such as cancer (as in the alterations in mitogenic pathways in melanoma). Hammell and colleagues approach this elemental problem by using a variety of model organisms and patient-derived cancer cell lines. To directly identify the components that function in controlling normal developmental timing, they use the small nematode Caenorhabditis elegans, applying forward and reverse genetic approaches. In contrast to the extreme robustness of cell-fate lineage in C. elegans, in which specification of developmental programs is hard-wired, mutations that alter conserved signaling pathways in melanoma create relatively plastic developmental landscapes that allow these lesions to become aggressive tumors. Notably, the gene regulatory architecture of melanoma cells allows them to acquire resistance to therapeutic agents. Hammell’s team is interested in epigenetic mechanisms that contribute to resistance, specifically dramatic changes in gene expression patterns and intracellular signaling pathways. They are performing high-throughput screens to identify cellular factors that allow these re-wiring events to occur, with the idea that these components would make ideal therapeutic targets to complement existing clinical strategies.
CSHL celebrates SBS Class of 2025
May 7, 2025
The CSHL School of Biological Sciences conferred nine Ph.D.s and one honorary Doctor of Science degree during its 22nd commencement ceremony.
At the Lab Season 1 Research Rewind: Genetics
October 22, 2024
It’s the code for all life on Earth. This week At the Lab, we’re hacking it with the help of Cold Spring Harbor Laboratory’s geneticists.
At the Lab Episode 10: The time of our lives
June 4, 2024
“You wouldn’t start making the fingernails on an arm until you had started to make the arm,” says CSHL’s Christopher Hammell. How’s that for a visual?
A quiz for the ages
January 29, 2024
Want to know the secret to a long life? So do CSHL scientists. Take this short quiz to see what they’ve found out about aging and longevity.
First look: Whole-body gene expression
November 6, 2023
For the first time, scientists at CSHL have observed gene expression as it occurs throughout an animal. See life take shape in front of your eyes.
These worms have rhythm
September 5, 2023
Observing gene expression in real time, CSHL scientists identified four molecules the C. elegans worm relies on to set the tempo of its development.
CSHL celebrates 20th graduating class
May 9, 2023
Friends, family, and faculty applauded 11 Ph.D. recipients at the CSHL School of Biological Sciences’ 2023 commencement.
CSHL 19th graduating class celebrated
May 2, 2022
In May 2022, the CSHL School of Biological Sciences awarded 10 Doctor of Philosophy degrees and two honorary degrees.
URP: Summer camp for undergrads
July 29, 2021
The Undergraduate Research Program brings college students from around the world to CSHL for a summer of research and fun.
How roundworms decide the time is right
December 22, 2020
Roundworm embryos calibrate the speed of their development to environmental conditions. A similar system may go wrong in some cancers.
All Publications
LIN-67 functionally interacts with heterochronic miRNAs and regulates developmental timing in Caenorhabditis elegans
2 Jan 2025 | bioRxiv
Medley, Jeffrey; Perales, Roberto; Humada, Belén; Doerfel, Katja; Levine, Ariana; Panzade, Ganesh; Hammell, Christopher; Zinovyeva, Anna;
Clr4SUV39H1 ubiquitination and non-coding RNA mediate transcriptional silencing of heterochromatin via Swi6 phase separation
30 Oct 2024 | Nature Communications | 15(1):9384
Kim, Hyun-Soo; Roche, Benjamin; Bhattacharjee, Sonali; Todeschini, Leila; Chang, An-Yun; Hammell, Christopher; Verdel, André; Martienssen, Robert;
Dynamic compartmentalization of the pro-invasive transcription factor NHR-67 reveals a role for Groucho in regulating a proliferative-invasive cellular switch in C. elegans
1 Dec 2023 | eLife | 12:RP84355
Medwig-Kinney, Taylor; Kinney, Brian; Martinez, Michael; Yee, Callista; Sirota, Sydney; Mullarkey, Angelina; Somineni, Neha; Hippler, Justin; Zhang, Wan; Shen, Kang; Hammell, Christopher; Pani, Ariel; Matus, David;
A circadian-like gene network programs the timing and dosage of heterochronic miRNA transcription during C. elegans development
22 Aug 2023 | Developmental Cell | :S1534-5807(23)00402
Kinney, Brian; Sahu, Shubham; Stec, Natalia; Hills-Muckey, Kelly; Adams, Dexter; Wang, Jing; Jaremko, Matt; Joshua-Tor, Leemor; Keil, Wolfgang; Hammell, Christopher;
An expandable FLP-ON::TIR1 system for precise spatiotemporal protein degradation in Caenorhabditis elegans
1 Feb 2023 | Genetics | :iyad013
Xiao, Yutong; Yee, Callista; Zhao, Chris; Martinez, Michael; Zhang, Wan; Shen, Kang; Matus, David; Hammell, Christopher;
An expandable FLP-ON::TIR1 system for precise spatiotemporal protein degradation in C. elegans
2022 | bioRxiv
Xiao, Yutong; Yee, Callista; Martinez, Michael; Zhao, Chris; Zhang, Wan; Shen, Kang; Matus, David; Hammell, Christopher;
PQN-59 antagonizes microRNA-mediated repression during post-embryonic temporal patterning and modulates translation and stress granule formation in C. elegans
Nov 2021 | PLoS Genetics | 17(11):e1009599
Carlston, Colleen; Weinmann, Robin; Stec, Natalia; Abbatemarco, Simona; Schwager, Francoise; Wang, Jing; Ouyang, Huiwu; Ewald, Collin; Gotta, Monica; Hammell, Christopher;
PQN-59 and GTBP-1 contribute to stress granule formation but are not essential for their assembly in C. elegans embryos
18 Oct 2021 | Journal of Cell Science
Abbatemarco, Simona; Bondaz, Alexandra; Schwager, Francoise; Wang, Jing; Hammell, Christopher; Gotta, Monica;
An engineered, orthogonal auxin analog/AtTIR1(F79G) pairing improves both specificity and efficacy of the auxin degradation system in Caenorhabditis elegans
12 Oct 2021 | Genetics
Hills-Muckey, Kelly; Martinez, Michael; Stec, Natalia; Hebbar, Shilpa; Saldanha, Joanne; Medwig-Kinney, Taylor; Moore, Frances; Ivanova, Maria; Morao, Ana; Ward, J; Moss, Eric; Ercan, Sevinc; Zinovyeva, Anna; Matus, David; Hammell, Christopher;
PQN-59 antagonizes microRNA-mediated repression and functions in stress granule formation during C. elegans development
2021 | bioRxiv
Carlston, Colleen; Weinmann, Robin; Stec, Natalia; Abbatemarco, Simona; Schwager, Francoise; Wang, Jing; Ouyang, Huiwu; Gotta, Monica; Hammell, Christopher;