Alea A. Mills
Professor
Cancer Center Member
Ph.D., University of California, Irvine, 1997
mills@cshl.edu | 516-367-6910
Cells employ stringent controls to ensure that genes are turned on and off at the correct time and place. Accurate gene expression relies on several levels of regulation, including how DNA and its associated molecules are packed together. I study the diseases arising from defects in these control systems, such as aging and cancer.
Alea Mills is studying genetic pathways important in cancer, aging, and autism, identifying the genetic players and determining how aberrations in their functions culminate in human disease. Through innovative use of a technique called “chromosome engineering,” the Mills group discovered that one of the most common genetic alterations in autism—deletion of a 27-gene cluster on chromosome 16—causes autism-like features in mice. These autism-like movement impairments can be identified just days after birth, suggesting that these features could be used to diagnose autism. Mills has also used chromosome engineering to identify a tumor suppressor gene that had eluded investigators for three decades. The gene, called Chd5, was shown by Mills to regulate an extensive cancer-preventing network. This year, the Mills lab uncovered how Chd5 acts as a tumor suppressor: It binds to a protein found within chromatin to turn specific genes on or off, halting cancer progression. The epigenetic role of Chd5 in development, cancer, and stem-cell maintenance is currently being investigated. The Mills lab is also studying p63 proteins, which regulate development, tumorigenesis, cellular senescence, and aging in vivo. They succeeded in halting the growth of malignant tumors by turning on production of one of the proteins encoded by the p63 gene, called TAp63. TAp63 also exerts other protective effects. This year, the Mills lab generated a mouse model which allowed them to find that TAp63 is required to prevent a genetic disorder, known as EEC (ectrodactyly-ectodermal dysplasia cleft lip/palate syndrome), which is characterized by a cleft palate and major deformities of the skin and limbs in infants. In addition, they recently discovered that a different version of p63, called ΔNp63, reprograms stem cells of the skin to cause carcinoma development—the most prevalent form of human cancer. Modulation of these proteins may offer new ways to treat human malignancies in the future.
Woman of the Year in Health/Medicine – 2012
‘The bridge between hope and healing’
June 4, 2025
The Cosmopolitan Club hosts an inspiring event featuring CSHL Professor Alea Mills and Dr. John Boockvar of Netflix’s Lenox Hill and Emergency NYC.
At the Lab Season 1 Research Rewind: Cancer
October 8, 2024
As the first season of our new podcast winds down, we’re revisiting all of our episodes with a focus on CSHL’s cutting-edge cancer research.
At the Lab Episode 24: Putting the brakes on brain cancer
September 17, 2024
CSHL Professor Alea Mills compares the deadly brain cancer glioblastoma to a car with its brakes cut. Her lab works to reattach them.
Pancreatic cancer’s cellular amnesia
June 17, 2024
New study from CSHL Professor Christopher Vakoc and former postdoc Diogo Maia-Silva shows how basal-like cancer cells lose their original identity.
The evolution of autism research
May 25, 2023
The conversation around autism has evolved over the past two decades. So has CSHL research. This retrospective shows how we’ve helped move the needle.
Cracking the mystery behind a deadly brain cancer
December 21, 2022
Scientists solve the mystery of how glioblastoma turns off cancer defenses without the usual cancer-inducing mutations.
Cold Spring Harbor Laboratory 2022 Ph.D.’s
May 1, 2022
The School of Biological Sciences awarded Ph.D. degrees to ten students this year. Here are some stories and memories from their time at CSHL.
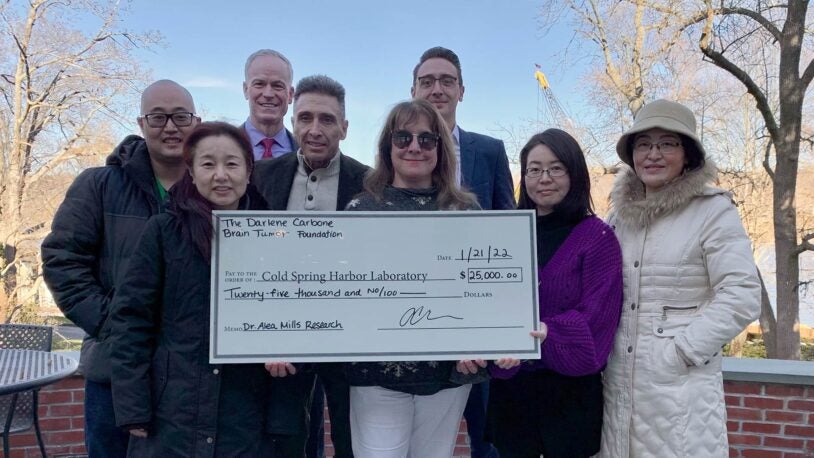
The Darlene Carbone Brain Tumor Foundation donates $25,000 to CSHL
January 24, 2022
The Darlene Carbone Brain Tumor Foundation donates $25,000 to Dr. Alea Mills lab for glioblastoma research.
Breaking the chain that culminates in cancer
December 1, 2021
CSHL scientists have discovered a way to shut down a cancer-causing protein by inhibiting a cascade of proteins that activate it.
CSHL graduate student wins HHMI fellowship
August 15, 2019
WSBS student David Johnson won a 2019 Gilliam Fellowships for Advanced Study for leadership and diversity in science.
All Publications
PRMT5/WDR77 Enhances the Proliferation of Squamous Cell Carcinoma via the ΔNp63α-p21 Axis
11 Nov 2024 | Cancers | 16(22):3789
Liang, Heng; Fisher, Matthew; Wu, Caizhi; Ballon, Carlos; Sun, Xueqin; Mills, Alea;
Cancer-associated fibroblasts promote cancer stemness by inducing expression of the chromatin-modifying protein CBX4 in squamous cell carcinoma
18 Aug 2023 | Carcinogenesis | 44(6):485-496
Fisher, Matthew; Balinth, Seamus; Hwangbo, Yon; Wu, Caizhi; Ballon, Carlos; Goldberg, Gary; Mills, Alea;
Chd5 regulates the transcription factor Six3 to promote neuronal differentiation
13 Jan 2023 | Stem Cells | :sxad002
Shrestha, Padmina; Jaganathan, Anbalagan; Huilgol, Dhananjay; Ballon, Carlos; Hwangbo, Yon; Mills, Alea;
BRD8 maintains glioblastoma by epigenetic reprogramming of the p53 network
21 Dec 2022 | Nature
Sun, Xueqin; Klingbeil, Olaf; Lu, Bin; Wu, Caizhi; Ballon, Carlos; Ouyang, Meng; Wu, Xiaoli; Jin, Ying; Hwangbo, Yon; Huang, Yu-Han; Somerville, Tim; Chang, Kenneth; Park, Jung; Chung, Taemoon; Lyons, Scott; Shi, Junwei; Vogel, Hannes; Schulder, Michael; Vakoc, Christopher; Mills, Alea;
ΔNp63α in cancer: importance and therapeutic opportunities
14 Sep 2022 | Trends in Cell Biology | :S0962-8924(22)00194
Fisher, Matthew; Balinth, Seamus; Mills, Alea;
EZH2 regulates a SETDB1/ΔNp63α axis via RUNX3 to drive a cancer stem cell phenotype in squamous cell carcinoma
21 Jul 2022 | Oncogene
Balinth, Seamus; Fisher, Matthew; Hwangbo, Yon; Wu, Caizhi; Ballon, Carlos; Sun, Xueqin; Mills, Alea;
P63 targeted deletion under the FOXN1 promoter disrupts pre-and post-natal thymus development, function and maintenance as well as induces severe hair loss
25 Jan 2022 | PLoS ONE | 17(1)
Stefanski, Heather; Xing, Yan; Nicholls, Jemma; Jonart, Leslie; Goren, Emily; Taylor, Patricia; Mills, Alea; Riddle, Megan; McGrath, John; Tolar, Jakub; Hollander, Georg; Blazar, Bruce;
BRD4 REGULATES TRANSCRIPTION FACTOR ∆Np63αTO DRIVE A CANCER STEM CELL PHENOTYPE IN SQUAMOUS CELL CARCINOMAS
25 Oct 2021 | Cancer Research | :canres.0707.2021
Fisher, Matthew; Balinth, Seamus; Hwangbo, Yon; Wu, Caizhi; Ballon, Carlos; Wilkinson, John; Goldberg, Gary; Mills, Alea;
p63-related signaling at a glance
11 Sep 2020 | Journal of Cell Science | 133(17)
Fisher, M; Balinth, S; Mills, A;
Reversal of Synaptic and Behavioral Deficits in a 16p11.2 Duplication Mouse Model via Restoration of the GABA Synapse Regulator Npas4
25 Feb 2020 | Molecular Psychiatry
Rein, B; Tan, T; Yang, F; Wang, W; Williams, J; Zhang, F; Mills, A; Yan, Z;