Christopher Vakoc
Professor
Alan and Edith Seligson Professor of Cancer Research
Cancer Center Deputy Director of Research
M.D., Ph.D., University of Pennsylvania, 2007
vakoc@cshl.edu | 516-367-5045
Cancer cells achieve their pathogenicity by changing which genes are on and off. To maintain these changes in gene expression, cancer cells rely on proteins that interact with DNA or modify chromatin. My group investigates how such factors sustain the aberrant capabilities of cancer cells, thereby identifying new therapeutic targets.
Cancer can be understood as a disease of dysfunctional gene expression control. Research in Chris Vakoc’s lab investigates how transcription factors and chromatin regulators cooperate to control gene expression and maintain the cancer cell state. This work makes extensive use of genetic screens to reveal cancer-specific functions for transcriptional regulators, as well as genomic and biochemical approaches to identify molecular mechanisms. One theme that has emerged from their efforts is that blood cancers are often vulnerable to targeting transcriptional coactivators, such as BRD4 and the SWI/SNF chromatin remodeling complex. Vakoc’s team demonstrated that chemical inhibition of BRD4 exhibits therapeutic effects in mouse models of leukemia, a finding that has motivated ongoing clinical trials in human leukemia patients. The Vakoc lab has also developed a CRISPR-Cas9 screening approach that can reveal individual protein domains that sustain cancer cells. Their lab is now deploying this technology in a diverse array of human cancers to reveal therapeutic opportunities and basic mechanisms of cancer gene control.
Vakoc wins Paul Marks Prize for cancer research
AACR Outstanding Achievement in Cancer Research Award
Forbeck Scholar Award
"V Scholar" by The V Foundation for Cancer Research
Burroughs Welcome Fund Career Award for Medical Scientists
Pershing Square Sohn Prize
Leukemia and Lymphoma Society Scholar Award
Powered by community, driven by science
February 26, 2026
CRF’s Angels Wish Gala fuels pediatric cancer research at Cold Spring Harbor Laboratory, advancing hope right before Rare Disease Month.
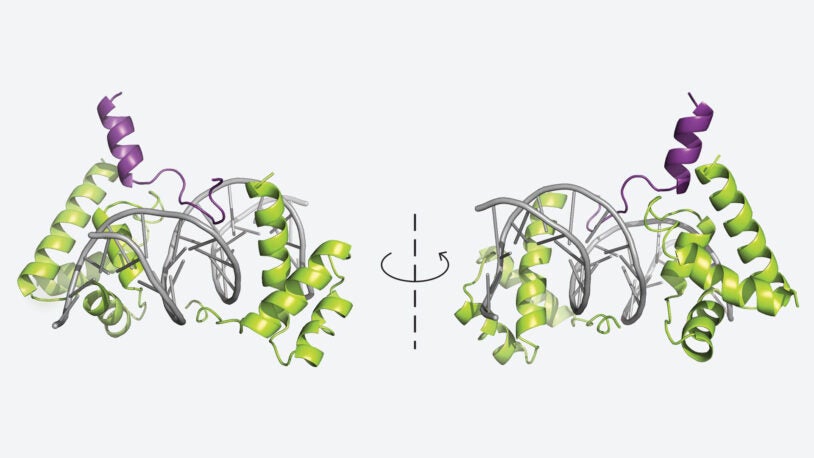
Shapeshifting cancers’ masters, unmasked
November 24, 2025
New research from CSHL’s Vakoc lab helps explain the bizarre behavior of certain pancreatic cancers and lung cancers that can change identities.
Pediatric cancers: Rare, relentless, and real
November 5, 2025
The science is complex. The stakes are high. And the urgency is immediate.
The CSHL School of Biological Sciences’ class of 2025
May 5, 2025
The School of Biological Sciences awarded Ph.D. degrees to nine students this year. Read some of their stories and reflections on their time at CSHL.
‘Her dream continues’
February 7, 2025
A night of remembrance, a generous donation, and another step closer to a cancer-free world.
Cocktails & Chromosomes: Cancer’s identity crisis
December 19, 2024
CSHL Professor Christopher Vakoc shares his research with a standing-room-only crowd at Industry bar in Huntington, NY.
Flipping cancer’s off switch
November 7, 2024
CSHL Professor Christopher Vakoc and postdoc Olaf Klingbeil have found a potential drug target for some of the most common and deadly cancers.
Friends of T.J. Foundation carries on his legacy
October 18, 2024
Since 2014, the organization has donated more than $500,000 in support of Cold Spring Harbor Laboratory’s cancer research.
At the Lab Season 1 Research Rewind: Cancer
October 8, 2024
As the first season of our new podcast winds down, we’re revisiting all of our episodes with a focus on CSHL’s cutting-edge cancer research.
At the Lab Episode 22: Outmuscling cancer
September 3, 2024
After 10 years, CSHL has made a breakthrough in the study of RMS, a rare pediatric cancer. How we got here is a story of innovation and perseverance.
All Publications
Systematic evaluation of GAPs and GEFs identifies a targetable dependency for hematopoietic malignancies
12 Aug 2025 | Cancer Discovery
Zhang, Pu; Cao, Zhendong; Pan, Xiangyu; Liu, Yuqiao; Castro, Cynthia; Kim, Won; Fujino, Takeshi; Lewis, Jennifer; Rahman, Jahan; Shahid, Sanam; Um, Jasmine; Burns, Erin; Chen, Bingyi; Cai, Winson; Ortiz-Pacheco, Juliana; Li, Zhuoning; Monetti, Mara; Vakoc, Christopher; Daniyan, Anthony; Abdel-Wahab, Omar; Shi, Junwei;
Correction: Targeting of epigenetic co-dependencies enhances anti-AML efficacy of Menin inhibitor in AML with MLL1-r or mutant NPM1.
21 May 2025 | Blood Cancer Journal | 15(1):99
Fiskus, Warren; Mill, Christopher; Birdwell, Christine; Davis, John; Das, Kaberi; Boettcher, Steffen; Kadia, Tapan; DiNardo, Courtney; Takahashi, Koichi; Loghavi, Sanam; Soth, Michael; Heffernan, Tim; McGeehan, Gerard; Ruan, Xinjia; Su, Xiaoping; Vakoc, Christopher; Daver, Naval; Bhalla, Kapil;
Histone chaperones coupled to DNA replication and transcription control divergent chromatin elements to maintain cell fate
16 Apr 2025 | Genes & Development
Franklin, Reuben; Zhang, Brian; Frazier, Jonah; Chen, Meijuan; Do, Brian; Padayao, Sally; Wu, Kun; Vander Heiden, Matthew; Vakoc, Christopher; Roe, Jae-Seok; Ninova, Maria; Murn, Jernej; Sykes, David; Cheloufi, Sihem;
PKM splice-switching ASOs induce upregulation of dual-specificity phosphatases and dephosphorylation of ERK1/2 in hepatocellular carcinoma
22 Feb 2025 | Journal of Biological Chemistry | :108345
Voss, Dillon; Kral, Alexander; Sim, GeunYoung; Utama, Raditya; Lin, Kuan-Ting; Cizmeciyan, Chris; Schafer, Balazs; Cunniff, Patrick; Vakoc, Christopher; Caruthers, Marvin; Mishra, Lopa; Krainer, Adrian;
Sequential epigenetic therapy in AML
13 Feb 2025 | Blood | 145(7):660-661
Wang, Yuanting; Vakoc, Christopher;
PTPN23-dependent ESCRT machinery functions as a cell death checkpoint
28 Nov 2024 | Nature Communications | 15(1):10364
Song, Dongyan; Cen, Yuxin; Qian, Zhe; Wu, Xiaoli; Rivera, Keith; Wee, Tse-Luen; Demerdash, Osama; Chang, Kenneth; Pappin, Darryl; Vakoc, Christopher; Tonks, Nicholas;
Dietary pro-oxidant therapy by a vitamin K precursor targets PI 3-kinase VPS34 function
25 Oct 2024 | Science | 386(6720):eadk9167
Swamynathan, Manojit; Kuang, Shan; Watrud, Kaitlin; Doherty, Mary; Gineste, Charlotte; Mathew, Grinu; Gong, Grace; Cox, Hilary; Cheng, Eileen; Reiss, David; Kendall, Jude; Ghosh, Diya; Reczek, Colleen; Zhao, Xiang; Herzka, Tali; Špokaitė, Saulė; Dessus, Antoine; Kim, Seung; Klingbeil, Olaf; Liu, Juan; Nowak, Dawid; Alsudani, Habeeb; Wee, Tse-Luen; Park, Youngkyu; Minicozzi, Francesca; Rivera, Keith; Almeida, Ana; Chang, Kenneth; Chakrabarty, Ram; Wilkinson, John; Gimotty, Phyllis; Diermeier, Sarah; Egeblad, Mikala; Vakoc, Christopher; Locasale, Jason; Chandel, Navdeep; Janowitz, Tobias; Hicks, James; Wigler, Michael; Pappin, Darryl; Williams, Roger; Cifani, Paolo; Tuveson, David; Laporte, Jocelyn; Trotman, Lloyd;
JMJD1C forms condensates to facilitate a RUNX1-dependent gene expression program shared by multiple types of AML cells
25 Oct 2024 | Protein and Cell | :pwae059
Chen, Qian; Wang, Saisai; Zhang, Juqing; Xie, Min; Lu, Bin; He, Jie; Zhen, Zhuoran; Li, Jing; Zhu, Jiajun; Li, Rong; Li, Pilong; Wang, Haifeng; Vakoc, Christopher; Roeder, Robert; Chen, Mo;
HDAC3 genetic and pharmacologic inhibition radiosensitizes fusion positive rhabdomyosarcoma by promoting DNA double-strand breaks
6 Aug 2024 | Cell Death Discovery | 10(1):351
Cassandri, Matteo; Porrazzo, Antonella; Pomella, Silvia; Noce, Beatrice; Zwergel, Clemens; Aiello, Francesca; Vulcano, Francesca; Milazzo, Luisa; Camero, Simona; Pajalunga, Deborah; Spada, Massimo; Manzi, Valeria; Gravina, Giovanni; Codenotti, Silvia; Piccione, Michela; Tomaciello, Miriam; Signore, Michele; Barillari, Giovanni; Marchese, Cinzia; Fanzani, Alessandro; De Angelis, Biagio; Quintarelli, Concetta; Vakoc, Christopher; Chen, Eleanor; Megiorni, Francesca; Locatelli, Franco; Valente, Sergio; Mai, Antonello; Rota, Rossella; Marampon, Francesco;
MARK2/MARK3 kinases are catalytic co-dependencies of YAP/TAZ in human cancer
26 Jul 2024 | Cancer Discovery
Klingbeil, Olaf; Skopelitis, Damianos; Tonelli, Claudia; Yoshimoto, Toyoki; Alpsoy, Aktan; Panepinto, Maria; Minicozzi, Francesca; Merrill, Joseph; Cafiero, Amanda; Aggarwal, Disha; Russo, Suzanne; Ha, Taehoon; Demerdash, Osama; Wee, Tse-Luen; Spector, David; Lyons, Scott; Tuveson, David; Cifani, Paolo; Vakoc, Christopher;