Hannah Meyer
Assistant Professor
Cancer Center Member
Ph.D., University of Cambridge, EMBL-EBI, 2018
hmeyer@cshl.edu | 516-367-8468
A properly functioning immune system must be able to recognize diseased cells and foreign invaders among the multitude of healthy cells in the body. This ability is essential to both prevent autoimmune diseases and fight infections and cancer. We study how a specific type of immune cells, known as T cells, are educated to make this distinction during development.
The thymus generates and selects a highly variable yet specific T cell repertoire which discriminates between healthy and non-healthy self and dangerous non-self antigens. My research group uses a systems immunology approach to dissect the mechanisms crucial to the selection processes in the thymus. We develop experimental techniques and combine the resulting data with innovative computational models to generate accurate and testable hypotheses about tissue-level organ physiology.
Studying thymus physiology from a qualitative and quantitative perspective will provide us with a more fine-grained understanding of the selection processes and their down-stream consequences such as auto-immunity, cancer immunosurveillance and immune deficiency.
The 2024 CSHL Volleyball Final
December 24, 2024
Eight teams entered the season on equal footing. Now, only two remain. But there can be only one champion. Press play to watch it all unfold.
Navlakha named Simons Foundation Pivot fellow
December 5, 2024
The computational biologist teams with CSHL’s Hannah Meyer to explore how the immune system solves problems also common in AI.
At the Lab Season 1 Research Rewind: AI+
October 29, 2024
This season’s final Research Rewind brings us from the realm of quantitative biology to neuroscience, genomics, and beyond.
The curious immune cells caught between worlds
September 24, 2024
CSHL’s Hannah Meyer shows innate-like T cells mature differently in humans and mice. Her discovery could improve preclinical immunotherapy studies.
The CSHL School of Biological Sciences’ class of 2024
May 5, 2024
The School of Biological Sciences awarded Ph.D. degrees to 11 students this year. Here are some stories and reflections from their time at CSHL.
At the Lab Episode 5: A heart of golf
April 30, 2024
A 500-year-old mystery stumbled on by Leonardo da Vinci has been solved using modern clinical data. Meet the CSHL scientist at the heart of it all.
The 2023 CSHL Volleyball League Finals
October 11, 2023
With a wide swath of the CSHL community in attendance, we got an up-close view of the action. How close? Think “camera on the ref’s head” close.
Eight serving one: CSHL volleyball mid-season report
August 2, 2023
CSHL’s 32nd Volleyball League season sees eight teams battling for the coveted Tiernan Cup and a year’s worth of bragging rights.
How popular steroids could mess up some cancer treatments
June 23, 2023
Scientists have long wondered how common steroids work and why cancer immunotherapy fails in certain patients. The answers may be one and the same.
President’s essay: Bringing bold visions to life
May 26, 2023
CSHL President & CEO Bruce Stillman sees the Laboratory as a global hub for scientific expertise and a powerful launchpad for early-career scientists.
All Publications
Unraveling the phenotypic states of human innate-like T cells: Comparative insights with conventional T cells and mouse models
10 Sep 2024 | Cell Reports | 43(9):114705
Loh, Liyen; Carcy, Salomé; Krovi, Harsha; Domenico, Joanne; Spengler, Andrea; Lin, Yong; Torres, Joshua; Prabakar, Rishvanth; Palmer, William; Norman, Paul; Stone, Matthew; Brunetti, Tonya; Meyer, Hannah; Gapin, Laurent;
BATMAN: Improved T cell receptor cross-reactivity prediction benchmarked on a comprehensive mutational scan database
25 Jan 2024 | bioRxiv
Banerjee, Amitava; Pattinson, David; Wincek, Cornelia; Bunk, Paul; Chapin, Sarah; Navlakha, Saket; Meyer, Hannah;
Unraveling the Phenotypic States of Human innate-like T Cells: Comparative Insights with Conventional T Cells and Mouse Models
8 Dec 2023 | bioRxiv
Loh, Liyen; Carcy, Salomé; Krovi, Harsha; Domenico, Joanne; Spengler, Andrea; Lin, Yong; Torres, Joshua; Palmer, William; Norman, Paul; Stone, Matthew; Brunetti, Tonya; Meyer, Hannah; Gapin, Laurent;
copepodTCR: Identification of Antigen-Specific T Cell Receptors with combinatorial peptide pooling.
29 Nov 2023 | bioRxiv
Kovaleva, Vasilisa; Pattinson, David; Barton, Carl; Chapin, Sarah; Minervina, Anastasia; Richards, Katherine; Sant, Andrea; Thomas, Paul; Pogorelyy, Mikhail; Meyer, Hannah;
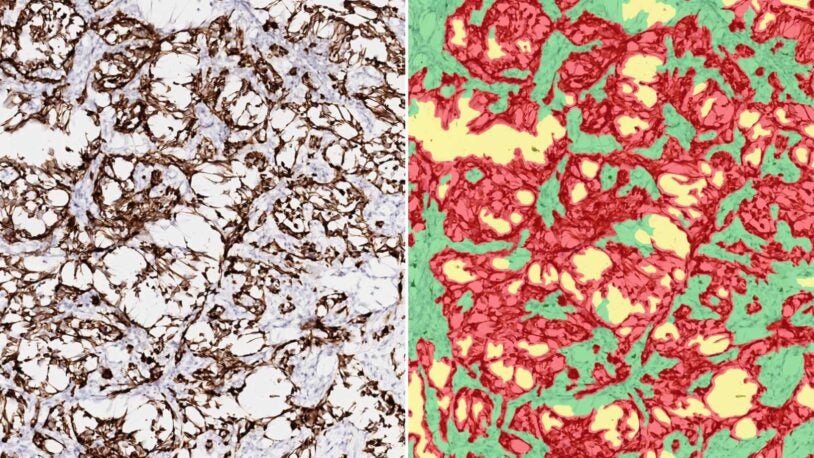
Cystatin C is glucocorticoid responsive, directs recruitment of Trem2+ macrophages, and predicts failure of cancer immunotherapy
9 Aug 2023 | Cell Genomics | 3(8):100347
Kleeman, Sam; Thakir, Tuba; Demestichas, Breanna; Mourikis, Nicholas; Loiero, Dominik; Ferrer, Miriam; Bankier, Sean; Riazat-Kesh, Yosef; Lee, Hassal; Chantzichristos, Dimitrios; Regan, Claire; Preall, Jonathan; Sinha, Sarthak; Rosin, Nicole; Yipp, Bryan; de Almeida, Luiz; Biernaskie, Jeff; Dufour, Antoine; Tober-Lau, Pinkus; Ruusalepp, Arno; Bjorkegren, Johan; Ralser, Markus; Kurth, Florian; Demichev, Vadim; Heywood, Todd; Gao, Qing; Johannsson, Gudmundur; Koelzer, Viktor; Walker, Brian; Meyer, Hannah; Janowitz, Tobias;
CRISPR-induced exon skipping of β-catenin reveals tumorigenic mutants driving distinct subtypes of liver cancer
15 Jan 2023 | Journal of Pathology
Mou, Haiwei; Eskiocak, Onur; Özler, Kadir; Gorman, Megan; Yue, Junjiayu; Jin, Ying; Wang, Zhikai; Gao, Ya; Janowitz, Tobias; Meyer, Hannah; Yu, Tianxiong; Wilkinson, John; Kucukural, Alper; Ozata, Deniz; Beyaz, Semir;
Snakeobjects: an object-oriented workflow management system
12 Dec 2022 | bioRxiv
Yamrom, Boris; Lee, Yoon-ha; Marks, Steven; Chorbadjiev, Lubomir; Meyer, Hannah; Iossifov, Ivan;
Cystatin C is associated with adverse COVID-19 outcomes in diverse populations
Oct 2022 | iScience | 25(10):105040
Kleeman, Sam; Cordioli, Mattia; Timmers, Paul; Khan, Atlas; Tober-Lau, Pinkus; Kurth, Florian; Demichev, Vadim; Meyer, Hannah; Wilson, James; Ralser, Markus; Kiryluk, Krzysztof; Ganna, Andrea; Baillie, Kenneth; Janowitz, Tobias;
Transcriptomic diversity in human medullary thymic epithelial cells
2 Aug 2022 | Nature Communications | 13(1):4296
Carter, Jason; Strömich, Léonie; Peacey, Matthew; Chapin, Sarah; Velten, Lars; Steinmetz, Lars; Brors, Benedikt; Pinto, Sheena; Meyer, Hannah;
Genetic and environmental determinants of diastolic heart function
13 Apr 2022 | Nature Cardiovascular Research | 1(4):361-371
Thanaj, Marjola; Mielke, Johanna; McGurk, Kathryn; Bai, Wenjia; Savioli, Nicolò; de Marvao, Antonio; Meyer, Hannah; Zeng, Lingyao; Sohler, Florian; Lumbers, R; Wilkins, Martin; Ware, James; Bender, Christian; Rueckert, Daniel; MacNamara, Aidan; Freitag, Daniel; O'Regan, Declan;