Adam Siepel
Professor
Cancer Center Member
Ph.D., University of California, Santa Cruz, 2005
asiepel@cshl.edu | 516-367-6922
I am a computer scientist who is fascinated by the challenge of making sense of vast quantities of genetic data. My research group focuses in particular on questions involving molecular evolution and transcriptional regulation, with applications to cancer and other diseases as well as to plant breeding and agriculture.
Modern genomic technologies make it relatively easy to generate rich data sets describing genome sequences, RNA expression, chromatin states, and many other aspects of the storage, transmission, and expression of genetic information. For many problems in genetics today, the limiting step is no longer in data generation, but in integrating, interpreting, and understanding the available data. Addressing these challenges requires expertise both in the practical arts of data analysis and in the theoretical underpinnings of statistics, computer science, and genetics.
My group focuses on a diverse collection of research questions in this interdisciplinary area, spanning applications in cancer biology, basic molecular biology, evolutionary genetics, infectious diseases, and agriculture. Over the years, our research has touched on topics including the identification of recombinant strains of HIV, the discovery of new human genes, the characterization of conserved regulatory elements in mammalian genomes, the identification of noncoding mutations important in cancer, and the discovery of ancient gene flow from humans to Neandertals. A general theme in our work is the development of precise mathematical models for the complex processes by which genomes evolve over time, and the use of these models, together with techniques from computer science and statistics, both to peer into the past, and to address questions of practical importance for human health. We collaborate closely with experimentalists in cancer biology, transcriptional regulation, plant breeding and many other areas.
John Simon Guggenheim Memorial Foundation Fellowship, 2012-2013
Alfred P. Sloan Foundation Research Fellowship, 2009-2011
David & Lucile Packard Foundation Fellowship for Science and Engineering, 2007
Microsoft Research Faculty Fellowship Program, 2007
National Science Foundation (NSF) CAREER Award, 2007
At the Lab Season 1 Research Rewind: AI+
October 29, 2024
This season’s final Research Rewind brings us from the realm of quantitative biology to neuroscience, genomics, and beyond.
How does cancer spread? Follow the map
September 25, 2024
CSHL Professor Adam Siepel and postdoc Armin Scheben use genetic barcodes to map how prostate cancer spreads.
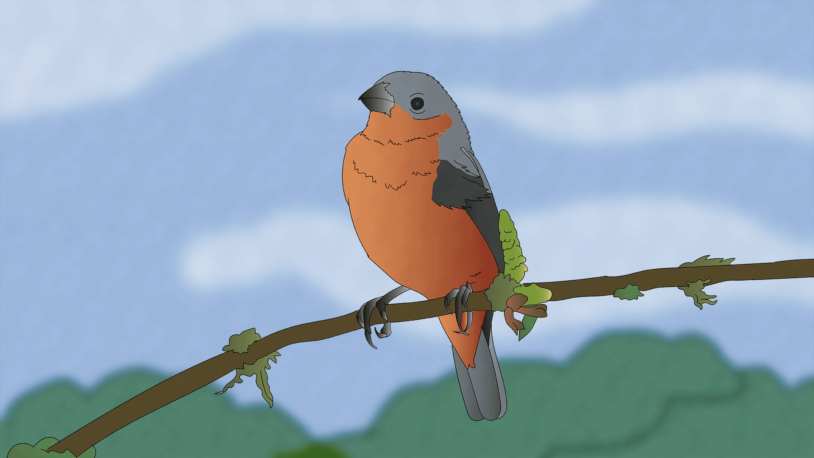
At the Lab Episode 8: Birds of a feather
May 21, 2024
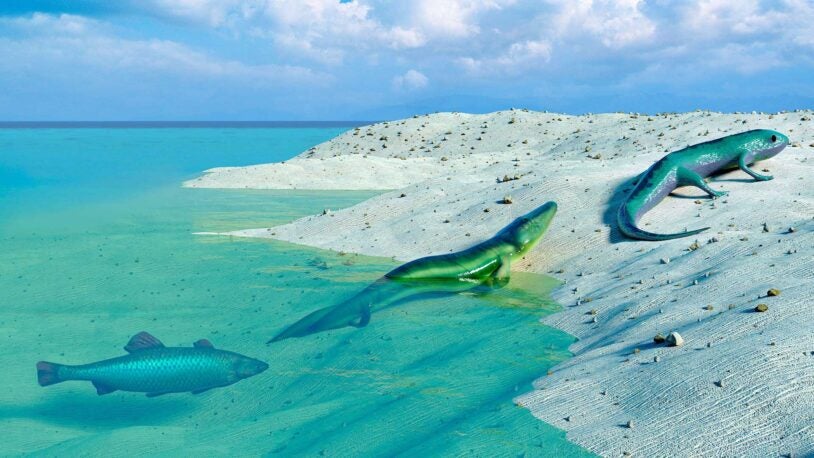
How did some birds get such distinct colors? CSHL Professor Adam Siepel joins us for a journey across evolution’s “islands of differentiation.”
The CSHL School of Biological Sciences’ class of 2024
May 5, 2024
The School of Biological Sciences awarded Ph.D. degrees to 11 students this year. Here are some stories and reflections from their time at CSHL.
A quiz for the ages
January 29, 2024
Want to know the secret to a long life? So do CSHL scientists. Take this short quiz to see what they’ve found out about aging and longevity.
Holy immunity! Bat genes key against COVID, cancer
October 16, 2023
Rapid evolution has streamlined bats’ immune systems. This may explain why they’re resistant to cancer and viruses like Ebola or COVID-19.
How evolved is your knowledge?
January 26, 2023
Test your knowledge of evolution with this quiz, inspired by the March 2023 performances of Isabella Rossellini’s play, Darwin’s Smile, at CSHL.
Exposing the evolutionary weak spots of the human genome
September 22, 2022
Researchers built a computer program that tracks harmful mutations throughout human evolution. It may help uncover the origins of genetic diseases.
CSHL Ph.D. program: Graduating class of 2021
August 22, 2021
The CSHL School of Biological Sciences awarded Ph.D. degrees to seven students this year, who describe some of their experiences.
Birds of a feather do flock together
November 17, 2020
Researchers found a genetic mechanism for how brand new species acquire distinct traits.
All Publications
A humanized NOVA1 splicing factor alters mouse vocal communications
18 Feb 2025 | Nature Communications | 16(1):1542
Tajima, Yoko; Vargas, César; Ito, Keiichi; Wang, Wei; Luo, Ji-Dung; Xing, Jiawei; Kuru, Nurdan; Machado, Luiz; Siepel, Adam; Carroll, Thomas; Jarvis, Erich; Darnell, Robert;
Probabilistic and machine-learning methods for predicting local rates of transcription elongation from nascent RNA sequencing data
8 Feb 2025 | Nucleic Acids Research (NAR) | 53(4)
Liu, Lingjie; Zhao, Yixin; Hassett, Rebecca; Toneyan, Shushan; Koo, Peter; Siepel, Adam;
Probabilistic and machine-learning methods for predicting local rates of transcription elongation from nascent RNA sequencing data.
8 Feb 2025 | Nucleic Acids Research (NAR) | 53(4)
Liu, Lingjie; Zhao, Yixin; Hassett, Rebecca; Toneyan, Shushan; Koo, Peter; Siepel, Adam;
Extensive genome evolution distinguishes maize within a stable tribe of grasses
24 Jan 2025 | bioRxiv
Stitzer, Michelle; Seetharam, Arun; Scheben, Armin; Hsu, Sheng-Kai; Schulz, Aimee; AuBuchon-Elder, Taylor; El-Walid, Mohamed; Ferebee, Taylor; Hale, Charles; La, Thuy; Liu, Zong-Yan; McMorrow, Sarah; Minx, Patrick; Phillips, Alyssa; Syring, Michael; Wrightsman, Travis; Zhai, Jingjing; Pasquet, Rémy; McAllister, Christine; Malcomber, Simon; Traiperm, Paweena; Layton, Daniel; Zhong, Jinshun; Costich, Denise; Dawe, R; Fengler, Kevin; Harris, Charlotte; Irelan, Zach; Llaca, Victor; Parakkal, Praveena; Zastrow-Hayes, Gina; Woodhouse, Margaret; Cannon, Ethalinda; Portwood, John; Andorf, Carson; Albert, Patrice; Birchler, James; Siepel, Adam; Ross-Ibarra, Jeffrey; Romay, M; Kellogg, Elizabeth; Buckler, Edward; Hufford, Matthew;
Teosinte Pollen Drive guides maize diversification and domestication by RNAi
7 Aug 2024 | Nature
Berube, Benjamin; Ernst, Evan; Cahn, Jonathan; Roche, Benjamin; de Santis Alves, Cristiane; Lynn, Jason; Scheben, Armin; Grimanelli, Daniel; Siepel, Adam; Ross-Ibarra, Jeffrey; Kermicle, Jerry; Martienssen, Robert;
Inferring polygenic negative selection underlying an individual trait as a distribution of fitness effects (DFEs) from GWAS summary statistics
2024 | bioRxiv
Xue, Alexander; Huang, Yi-Fei; Siepel, Adam;
Clonal Lineage Tracing with Somatic Delivery of Recordable Barcodes Reveals Migration Histories of Metastatic Prostate Cancer
5 Jul 2024 | Cancer Discovery
Serio, Ryan; Scheben, Armin; Lu, Billy; Gargiulo, Domenic; Patruno, Lucrezia; Buckholtz, Caroline; Chaffee, Ryan; Jibilian, Megan; Persaud, Steven; Staklinski, Stephen; Hassett, Rebecca; Brault, Lise; Ramazzotti, Daniele; Barbieri, Christopher; Siepel, Adam; Nowak, Dawid;
Single-Cell Transcription Mapping of Murine and Human Mammary Organoids Responses to Female Hormones
30 Jan 2024 | Journal of Mammary Gland Biology and Neoplasia | 29(1):3
Ortiz, Jenelys; Lewis, Steven; Ciccone, Michael; Chatterjee, Deeptiman; Henry, Samantha; Siepel, Adam; Dos Santos, Camila;
DNA-sequence and epigenomic determinants of local rates of transcription elongation
23 Dec 2023 | bioRxiv
Liu, Lingjie; Zhao, Yixin; Siepel, Adam;
Identification of constrained sequence elements across 239 primate genomes
29 Nov 2023 | Nature
Kuderna, Lukas; Ulirsch, Jacob; Rashid, Sabrina; Ameen, Mohamed; Sundaram, Laksshman; Hickey, Glenn; Cox, Anthony; Gao, Hong; Kumar, Arvind; Aguet, Francois; Christmas, Matthew; Clawson, Hiram; Haeussler, Maximilian; Janiak, Mareike; Kuhlwilm, Martin; Orkin, Joseph; Bataillon, Thomas; Manu, Shivakumara; Valenzuela, Alejandro; Bergman, Juraj; Rouselle, Marjolaine; Silva, Felipe; Agueda, Lidia; Blanc, Julie; Gut, Marta; de Vries, Dorien; Goodhead, Ian; Harris, R; Raveendran, Muthuswamy; Jensen, Axel; Chuma, Idriss; Horvath, Julie; Hvilsom, Christina; Juan, David; Frandsen, Peter; Schraiber, Joshua; de Melo, Fabiano; Bertuol, Fabrício; Byrne, Hazel; Sampaio, Iracilda; Farias, Izeni; Valsecchi, João; Messias, Malu; da Silva, Maria; Trivedi, Mihir; Rossi, Rogerio; Hrbek, Tomas; Andriaholinirina, Nicole; Rabarivola, Clément; Zaramody, Alphonse; Jolly, Clifford; Phillips-Conroy, Jane; Wilkerson, Gregory; Abee, Christian; Simmons, Joe; Fernandez-Duque, Eduardo; Kanthaswamy, Sree; Shiferaw, Fekadu; Wu, Dongdong; Zhou, Long; Shao, Yong; Zhang, Guojie; Keyyu, Julius; Knauf, Sascha; Le, Minh; Lizano, Esther; Merker, Stefan; Navarro, Arcadi; Nadler, Tilo; Khor, Chiea; Lee, Jessica; Tan, Patrick; Lim, Weng; Kitchener, Andrew; Zinner, Dietmar; Gut, Ivo; Melin, Amanda; Guschanski, Katerina; Schierup, Mikkel; Beck, Robin; Karakikes, Ioannis; Wang, Kevin; Umapathy, Govindhaswamy; Roos, Christian; Boubli, Jean; Siepel, Adam; Kundaje, Anshul; Paten, Benedict; Lindblad-Toh, Kerstin; Rogers, Jeffrey; Marques Bonet, Tomas; Farh, Kyle;