Plant Biology
 Plant research at CSHL explores fundamental mechanisms in plant development and genetics with a goal of increasing crop productivity and biodiversity, and reducing climate change through exploring the potential of biofuels.
Plant research at CSHL explores fundamental mechanisms in plant development and genetics with a goal of increasing crop productivity and biodiversity, and reducing climate change through exploring the potential of biofuels.
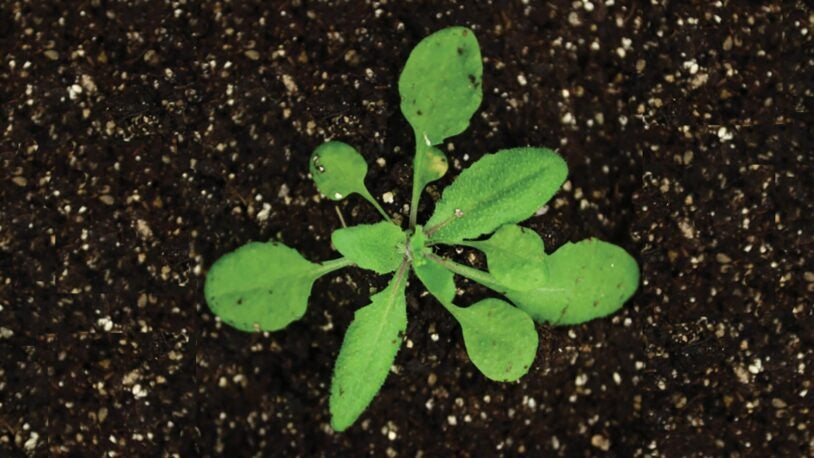
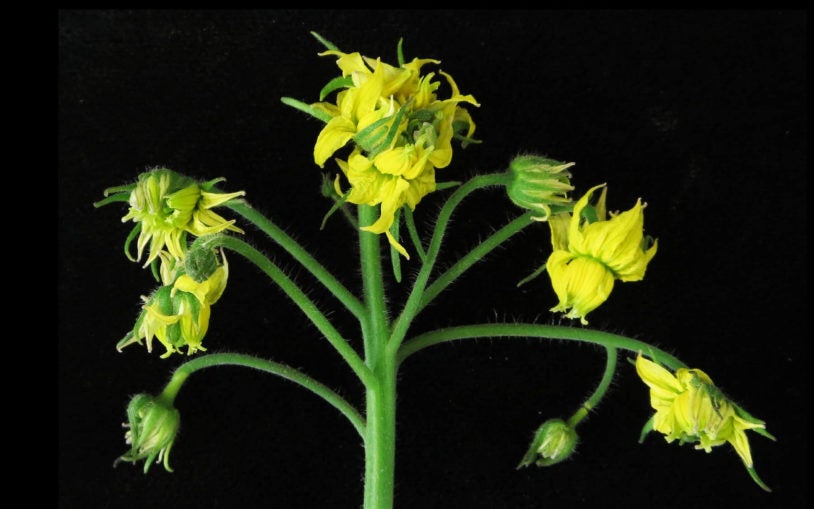
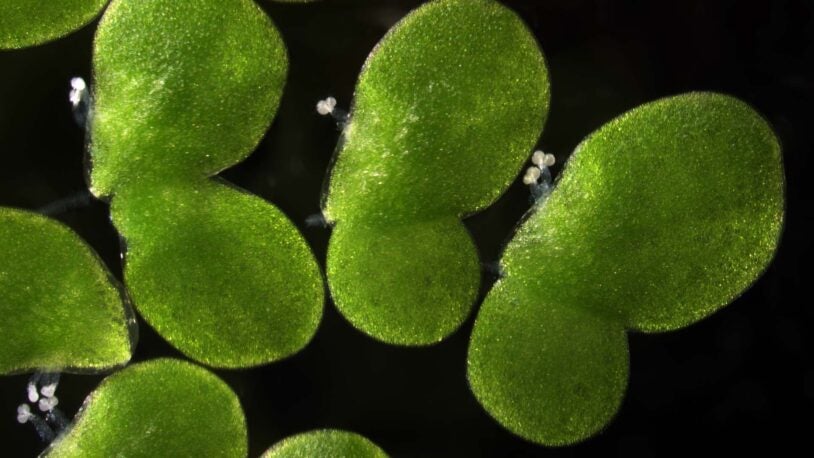
The plant biology group at CSHL focuses on plant development and gene expression, in an effort to uncover basic mechanisms that could lead to increased crop productivity, increased biodiversity and exploring the potential of biofuels. Researchers use Arabidopsis, maize, tomato and duckweed as model systems to uncover the principles that govern plant growth. Much of this work takes place on 12 acres of farmland at the nearby CSHL Uplands Farm, where expert staff raise crops and Arabidopsis plants for study. Research also involves bioinformatics and quantitative analysis of large data sets for functional genomics and developmental genetics, and has contributed to more than two dozen large scale collaborative genome projects funded by the National Science Foundation, the Department of Energy, and the United States Department of Agriculture.
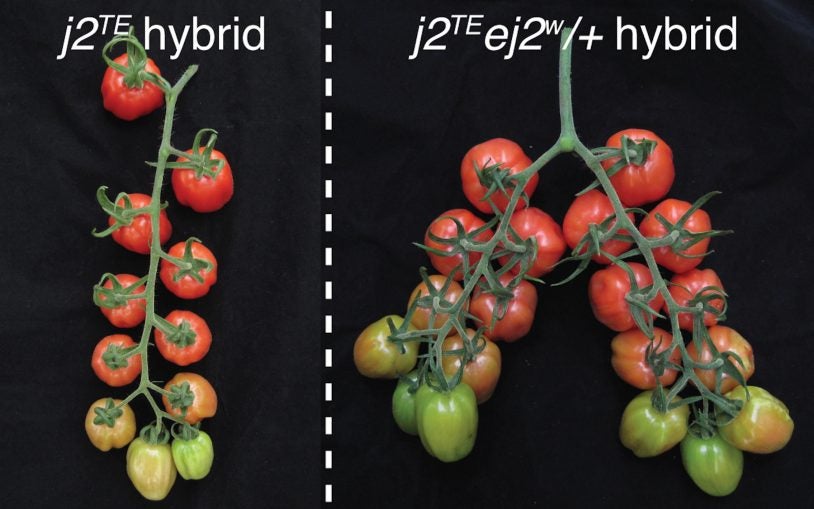
At CSHL, plant research has a storied history, including Nobel prize-winning research done by Barbara McClintock in the 1940s and 50s. The transposable genetic elements, or “jumping genes,” that she discovered decades ago are now understood to reprogram the epigenome, and are used as research tools by current CSHL researchers studying plant genomes.
Environmental science’s Cold Spring Harbor origin
April 22, 2024
In 1929, Ruth Patrick came to CSHL to study plant life. She’d meet her future husband here and go on to pioneer an entirely new field of biology.
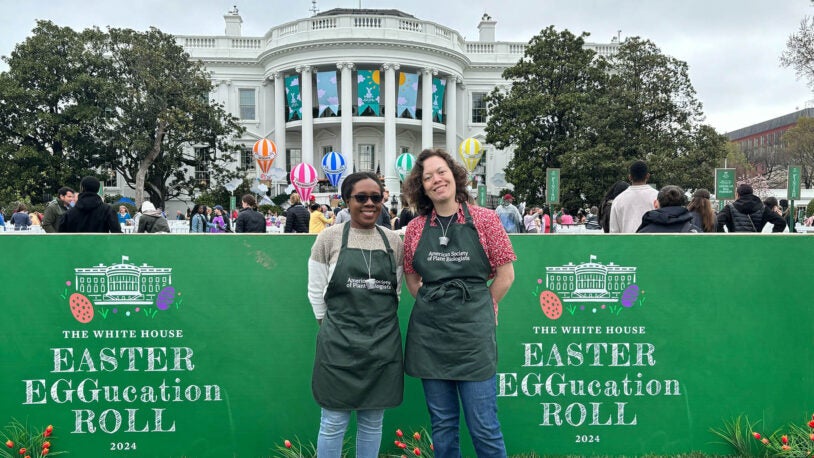
CSHL goes to the White House for Easter EGGucation
April 8, 2024
CSHL plant scientists taught kids the basics of plant biology and its role in the environment at an event hosted by First Lady Jill Biden.
CSHL’s Thomas Gingeras awarded $2 million NSF grant
April 3, 2024
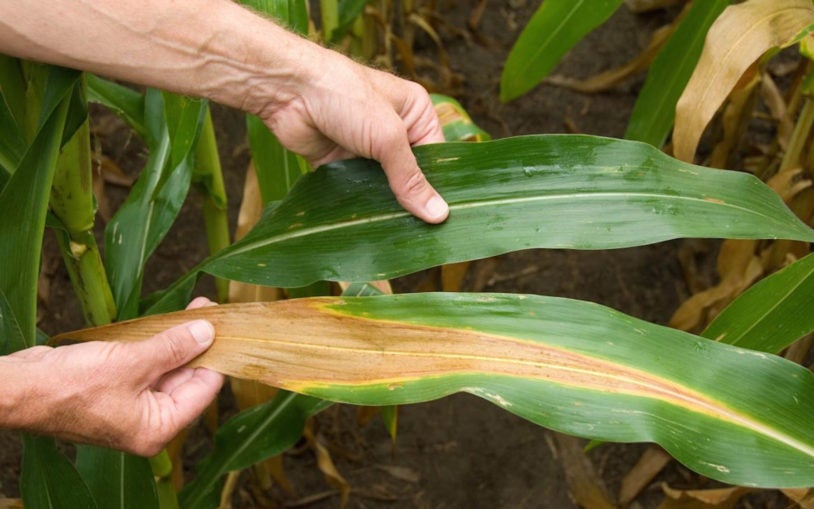
Climate change threatens crops with acidic soils and aluminum toxicity. Gingeras leads an international team tackling this problem head-on.
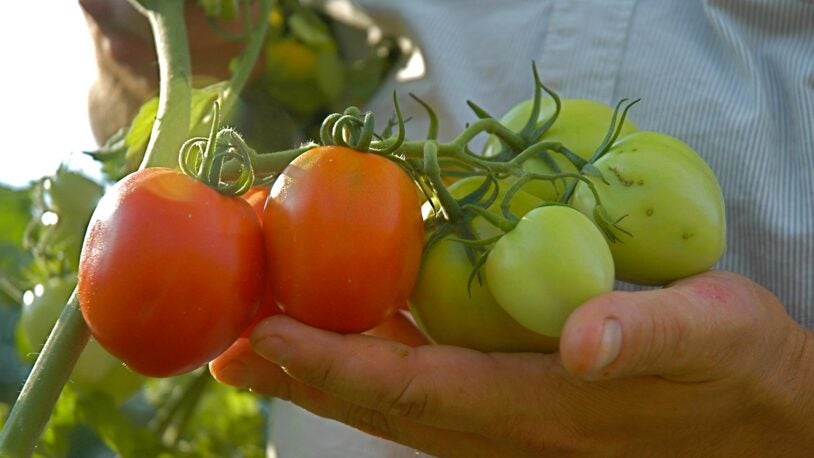
At the Lab Episode 1: The tomato of tomorrow
April 2, 2024
In the premiere episode of At the Lab, we visit CSHL Professor & HHMI Investigator Zachary Lippman to glimpse the future of food and farming.
From plant genomics to a bioscience revolution
March 25, 2024
CSHL played a lead role in mapping the first plant genome. Today, that breakthrough fuels a whole new understanding of life on Earth.
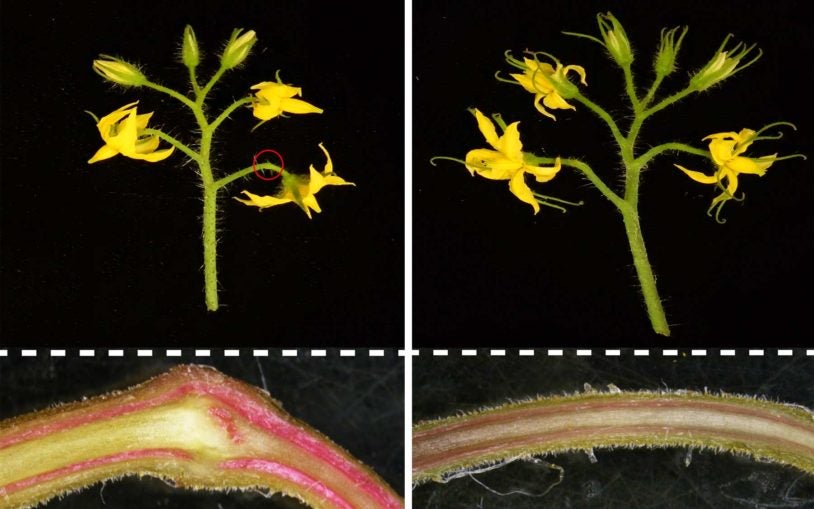
An evolutionary mystery 125 million years in the making
March 4, 2024
CSHL plant biologists have stumbled on a peculiar case involving a gene that’s key for controlling growth in tomatoes and other crops.
Cocktails & Chromosomes: Best shots of 2023
January 16, 2024
After a four-year hiatus, Cocktails & Chromosomes returned in 2023. Relive the past year’s best moments and see what’s in store for 2024.
Joshua-Tor named CSHL Director of Research
January 2, 2024
The Cold Spring Harbor Laboratory professor and HHMI investigator steps into her new role effective January 2, 2024.
Rob Martienssen awarded 2024 Genetics Society Medal
November 8, 2023
CSHL Professor Rob Martienssen earned the award for outstanding research contributions to the field of genetics.
You say genome editing, I say natural mutation
October 19, 2023
CSHL scientists have discovered that evolution and genome editing in crops are less predictable than previously thought.
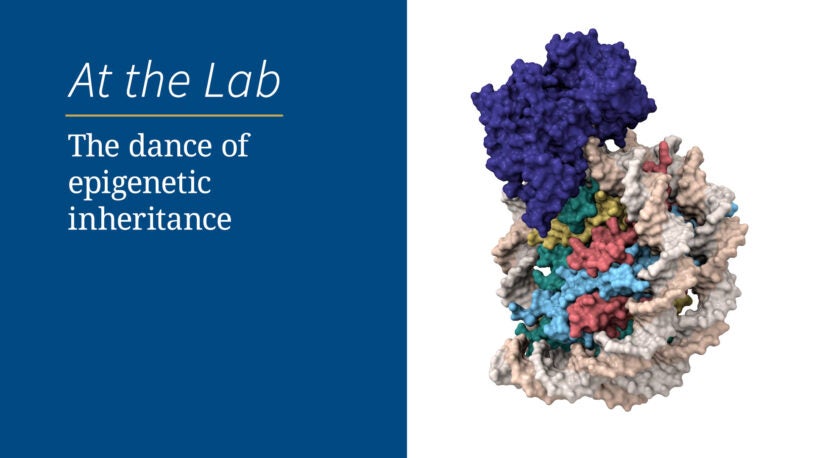
The dance of epigenetic inheritance
September 27, 2023
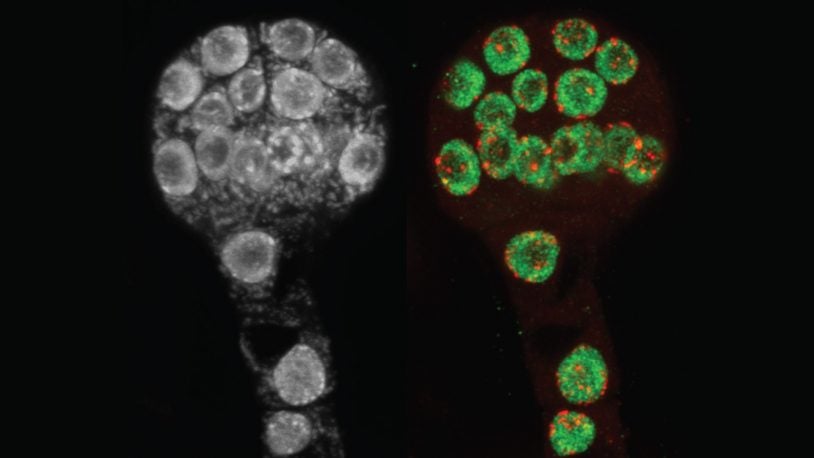
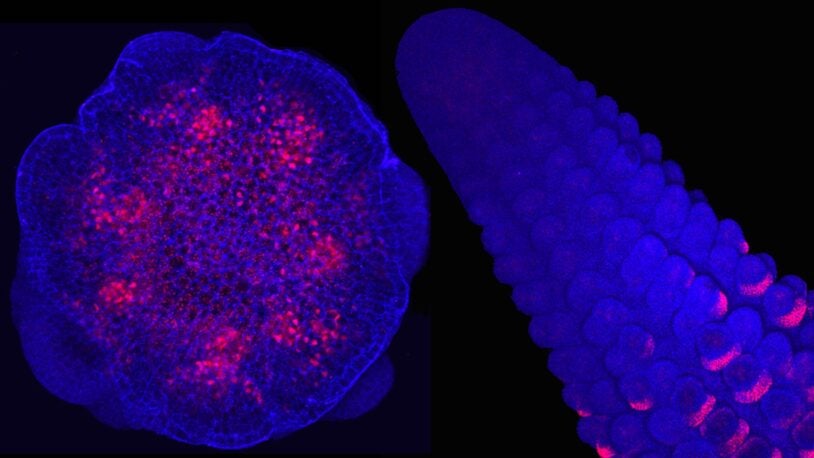
Making sure chromosomes get passed down correctly is hard work. Watch, through fluorescent and cryogenic lenses, how two proteins make it happen.
Cocktails & Chromosomes: Uprooting climate change
September 21, 2023
Pull up a stool to watch CSHL Associate Professor Ullas Pedmale speak about plants and climate change at Industry bar in Huntington, NY.
How plants pass down genetic memories
August 28, 2023
Thirty years ago, CSHL’s Rob Martienssen discovered plant gene DDM1. Now, he’s identified just how the DDM1 protein helps control inheritance.
Cocktails & Chromosomes gets corny
August 10, 2023
In the best way possible! CSHL Professor David Jackson talked about corn genetics. Plus, we gave away tickets to Tony Award-winning musical Shucked.
CSHL’s Cocktails & Chromosomes series returns this summer
June 22, 2023
The event, serving up stimulating science talks over delicious drinks, heads to Industry bar in Huntington, New York. First on tap for June 29 is AI.
Climate-control technology future-proofs plants
June 13, 2023
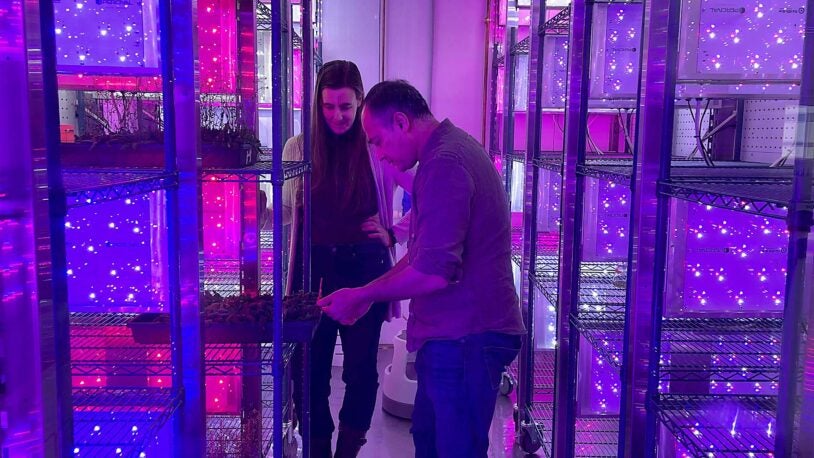
Take a virtual tour of CSHL’s new state-of-the-art growth chambers with plant biologist Ullas Pedmale.
New CSHL technology controls the weather and time itself
June 8, 2023
State-of-the-art plant growth chambers at CSHL allow scientists to mimic the effects of climate change on crops around the world.
Digging up the deep-rooted secrets of perennial corn
March 10, 2023
Perennials may hold the key to sustainable farming. CSHL scientists are decoding the genes that let these plants withstand the test of time.
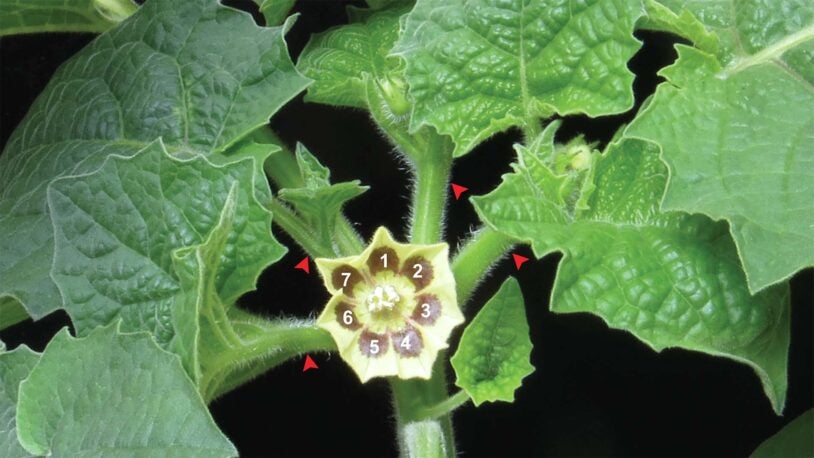
Tomato cousin’s coolest quirk
January 25, 2023
Take an up-close look at what CSHL Professor Zachary Lippman describes as “one of the coolest evolutionary novelties to emerge in plants.”
Reinvigorating CSHL’s Ph.D. program
December 13, 2022
Graduate student turned director of graduate studies, Zachary Lippman shares his vision for the CSHL School of Biological Sciences.
The tiny plant tackling climate change
December 8, 2022
The humble aquatic duckweed plant has enormous potential as a new source of healthy protein, low-carbon biofuels, and other bioproducts.
Welcome to Biology + Beyond
November 14, 2022
CSHL President and CEO Bruce Stillman introduces a special issue of Nautilus magazine now online, featuring the Lab’s latest groundbreaking research
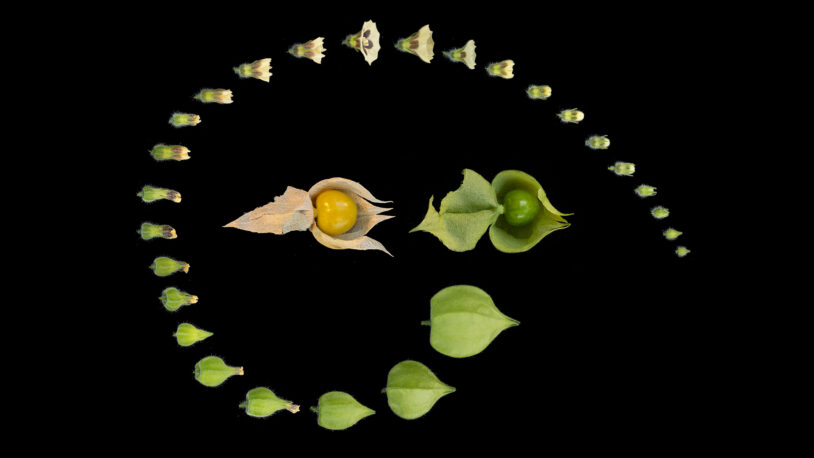
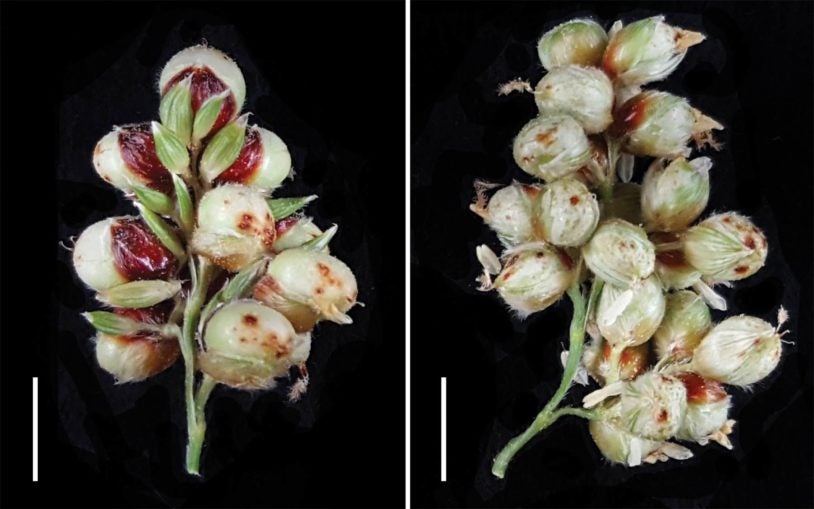
CSHL groundcherry research bears new fruits
October 31, 2022
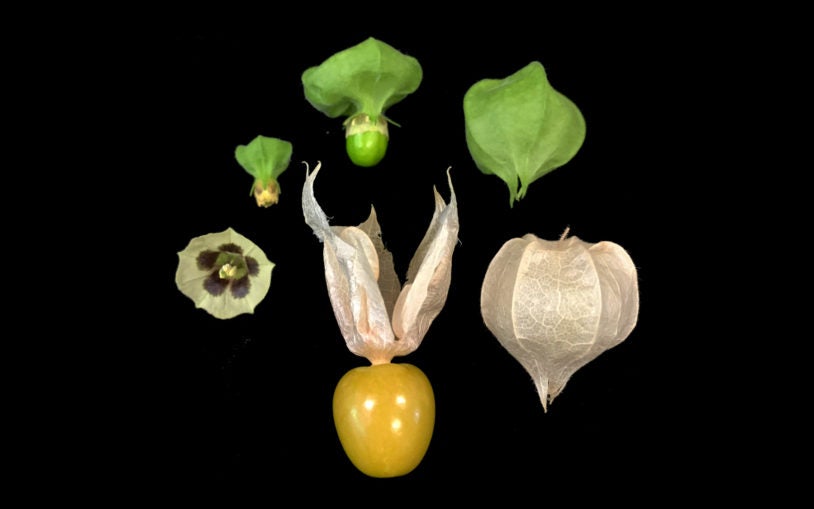
New genetic blueprints for two types of groundcherry may help strengthen food supplies and reveal how plants evolve.
What shedding light on plant growth could mean for cancer
June 13, 2022
CSHL researchers have found a new way to control plant growth by manipulating proteins involved in the process of detecting light.
How tweaking genes keeps corn and rice on your plate
May 18, 2022
CSHL scientists leveraged lessons from the corn genome to improve another global staple crop, rice.
Martienssen elected to American Academy of Arts and Sciences
April 28, 2022
CSHL Professor and HHMI Investigator Rob Martienssen joins the American Academy of Arts and Sciences.
Do you have the dirt on plant research?
March 31, 2022
New research is constantly sprouting. Take this quiz and test your plant knowledge.
For plant geneticists, some genes are double the trouble
March 28, 2022
It pays to check whether genetic tweaks that improve one kind of crop could get foiled by backup genes in a different crop.
Stabilizing chromosomes to tackle tumors
March 17, 2022
Dicer and its partner BRD4 stabilize chromosomes. Targeting this pair could provide new therapeutic opportunities against cancer.
Plants fight for their lives
January 31, 2022
As arable land disappears, a genetic tweak might secure the world’s food supply.
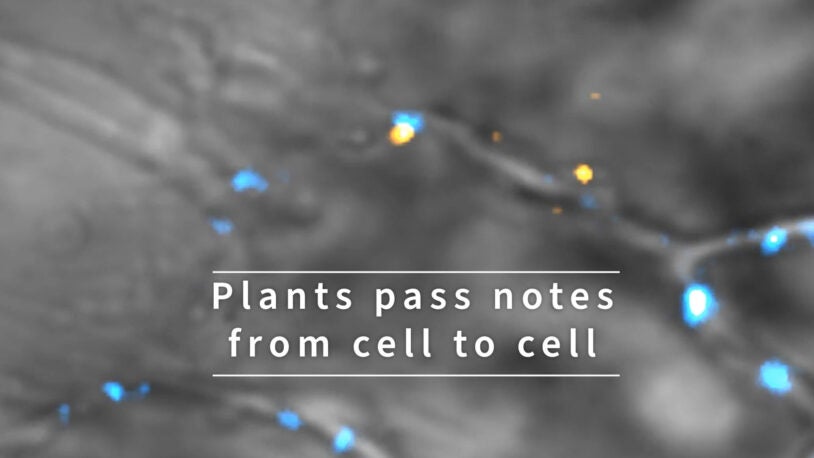
Psst! Plants pass notes from cell to cell
January 14, 2022
Plants relay messages critical for development between their cells using RNA. See tiny RNA messages zoom around inside plant cells.
Plants: RNA notes to self
January 13, 2022
CSHL scientists discovered a way plants send messages between cells using RNA and a protein escort.
Lab life: A step for students, a leap for science
December 20, 2021
The CSHL Partners for the Future program allows high school students to participate in laboratory life.
In the Field: A Barbara McClintock–inspired novel
December 2, 2021
1983 Nobel laureate Barbara McClintock continues to inspire many today. In 2021, author Rachel Pastan published a novel based on her life and legacy.
Corn 2.0
November 29, 2021
Climate change and population growth are threatening our crops. CSHL Professor David Jackson is helping corn keep up with the demand.
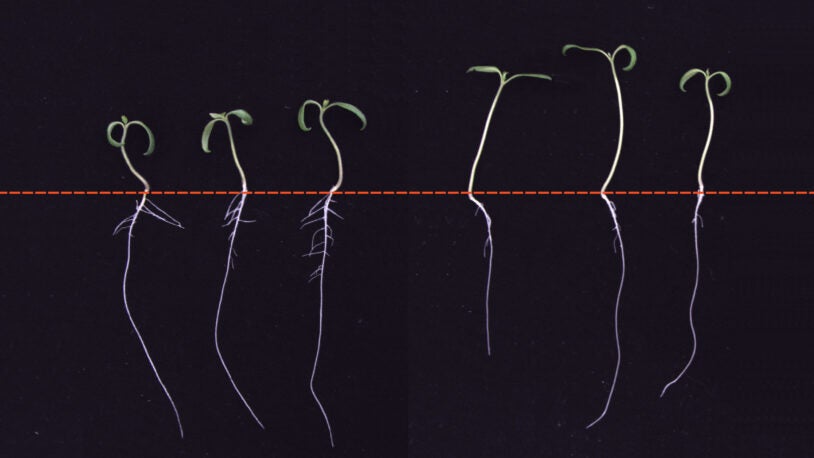
Why roots don’t grow in the shade
October 27, 2021
CSHL researchers found that sun-loving plants grown in the shade express stress hormones, which stunt their root systems.
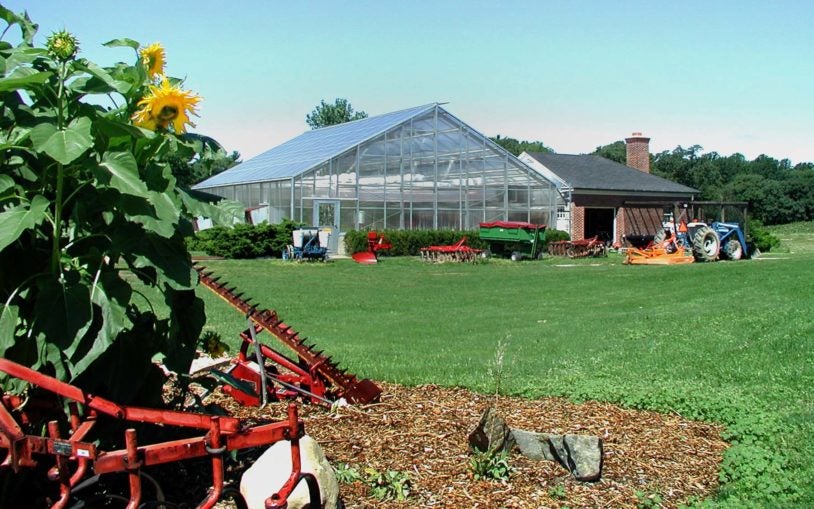
Tools of the trade at CSHL: The greenhouse
September 13, 2021
The greenhouse provides a beautiful space for hundreds of experimental plants to grow all year round, which could allow scientists to build better crops so
Using “guilt by association” to classify cells
July 14, 2021
Using a new computational statistics tool, CSHL researchers classify cells to understand how an organism functions.
How plants leave behind their parents’ genomic baggage
May 20, 2021
A baby plant resets its genome, erasing the changes that its parents accumulated. CSHL scientists found how the plant adds back a few necessary ones.
For tomato genes, one plus one doesn’t always make two
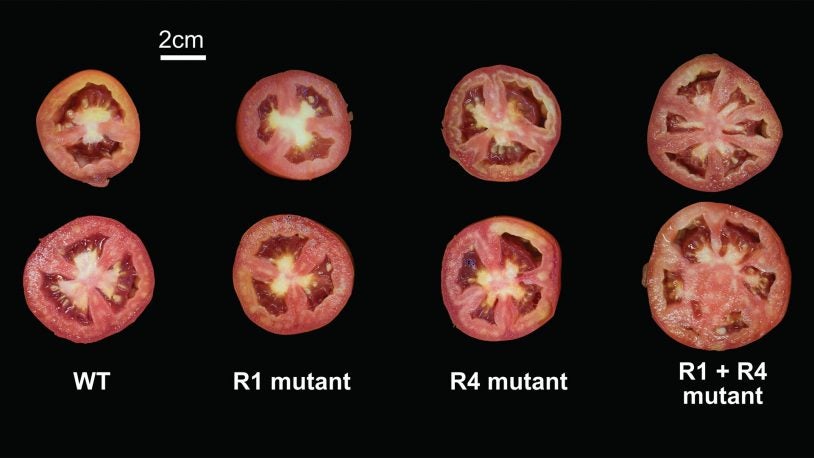
April 12, 2021
CSHL scientists made a series of mutations in tomatoes to see how they interact with one other. Some combinations yield larger changes than expected.
WOX9: A jack of all trades
March 4, 2021
A single gene can have many functions across different plant genomes. Changing the gene’s regulatory region can change the traits it produces.
Tweaking corn kernels with CRISPR
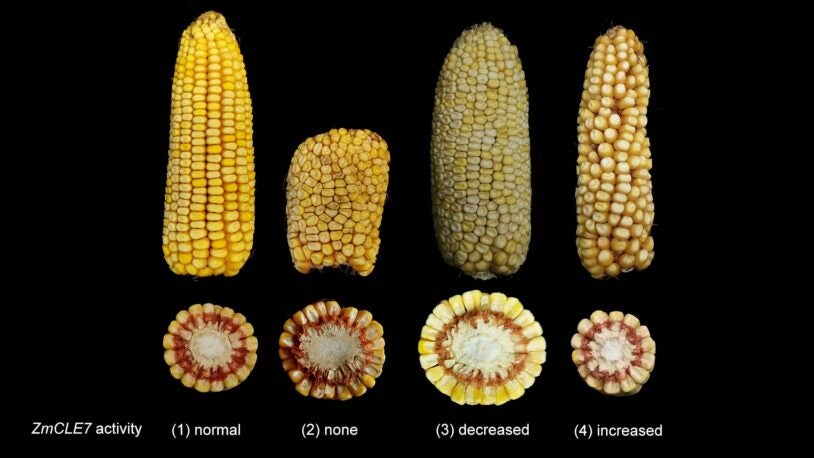
February 22, 2021
CRISPR genome editing can fast-forward the process of plant evolution. Researchers at CSHL are using the technique to increase kernel yield.
A healthy use for tobacco in coronavirus research
February 11, 2021
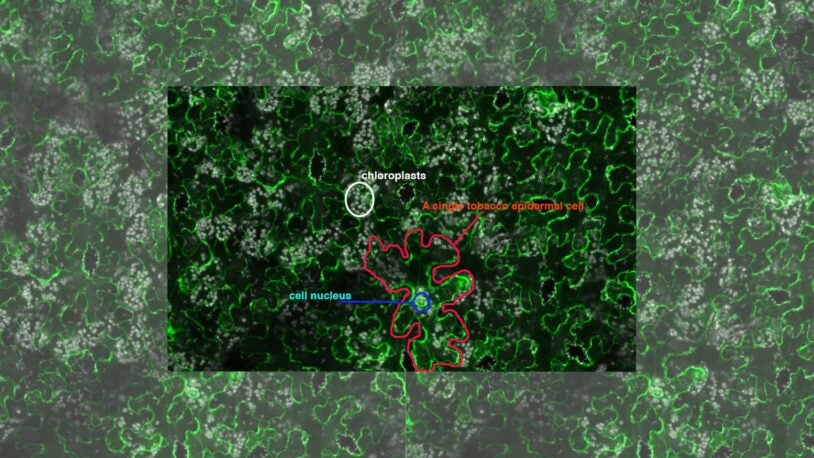
CSHL plant scientists grew fragments of coronavirus proteins in tobacco. They hoped to provide a cheap source of protein for virus and vaccine researchers.
How to bury carbon? Let plants do the dirty work.
February 5, 2021
Carbon sequestration could slow or reverse human emissions—and nothing is better at sequestration than a green plant.
Building a corn cob—cell by cell, gene by gene
January 26, 2021
CSHL scientists are piecing together the genes that control how corn develops.
Designing crops for a changing climate
December 15, 2020
In the face of climate change, engineering genetic modifications into new crops will ensure future food supplies.
Leshan, Lippman address “Life Science Across the Globe” series
September 11, 2020
Professor Zach Lippman and Executive Director of the Banbury Center Rebecca Leshan give talks as part of the “Life Science Across the Globe” series.
Human Nature documentary panel discussion
September 2, 2020
CRISPR experts, including Professors Zach Lippman, Jennifer Doudna, and Alta Charo discuss the gene-editing technology and its use.
CSHL hosts Human Nature documentary panel with CRISPR experts
September 2, 2020
CRISPR experts, including Professors Zach Lippman, Jennifer Doudna, and Alta Charo discuss the gene-editing technology and its use.
Martienssen named 2020 Royal Society winner
August 3, 2020
Professor and HHMI Investigator Rob Martienssen wins a 2020 Royal Society medal for his RNAi research.
The world destroyer in your shampoo and ice cream
July 6, 2020
Palm oil is an environmental scourge. Genetics has a solution.
Nobelist Sir Richard Roberts talks GMOs at CSHL
June 26, 2020
Nobelist and CSHL alum Sir Richard Roberts spoke about GMOs and the future of agriculture with Pamela Ronald and Rob Martienssen.
Nobelist Sir Richard Roberts talks GMOs at CSHL hosted event
June 25, 2020
Nobelist and CSHL alum Sir Richard Roberts spoke about GMOs and the future of agriculture with Pamela Ronald and Rob Martienssen in this video.
The DNA tricks that gave us 100 different kinds of tomatoes
June 17, 2020
It takes 230,000 genetic differences to make 100 different varieties of tomatoes.
Coronavirus research in plants
May 15, 2020
Purified coronavirus proteins are in short supply for COVID-19 researchers, so CSHL plant scientists are jumping in to make them.
Zachary Lippman wins Charles Albert Shull Award
April 8, 2020
Professor Zachary Lippman is the 2020 Charles Albert Shull Award winner, an honor given by the American Society for Plant Biology.
The future of food looks small, dense, and very bushy
February 18, 2020
Vertical farming could make agriculture more robust and sustainable. To unlock that potential, scientists are redesigning crops for urban life.
Lippman wins NAS Prize in Food and Agriculture Sciences
January 22, 2020
CSHL Professor and HHMI Investigator Zachary Lippman was awarded the NAS Prize for his work in the field of plant genetics.
A new tomato ideal for urban gardens and even outer space
December 23, 2019
Researchers used CRISPR gene editing to optimize tomatoes for urban agriculture, preparing them for the city rooftops and possibly space missions.
Shifting the balance of growth vs. defense boosts crop yield
December 19, 2019
Researchers found that a specific gene in maize balances both growth of the plant and its immunity.
Plant scientist David Jackson–A CSHL PI profile
November 5, 2019
CSHL Professor David Jackson studies mutated corn and flowers.
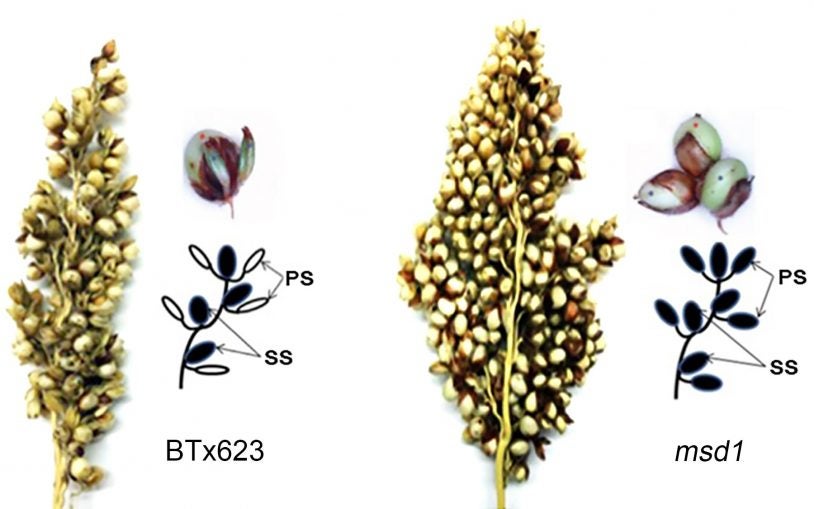
Researchers double sorghum grain number to improve food supply
October 30, 2019
A set of hormone-controlling genes may be the key to doubling grain number in sorghum plants.
The important role of an Uplands Farm cat
October 29, 2019
Cats, Uplands Farm’s smallest employees, have always played a big role in plant care and seed guarding.
Zachary Lippman named 2019 MacArthur Fellow
September 25, 2019
Cold Spring Harbor Laboratory Professor and HHMI Investigator Zachary Lippman has been named a 2019 MacArthur Fellow.
The next agricultural revolution is here
September 19, 2019
After reviewing decades of plant research, scientists suggest that with past lessons and modern tools, the next agricultural revolution is at hand.
Engineering Plants for Agriculture
September 11, 2019
Examines the molecular bases of plant traits and addresses how this knowledge can be used to develop crops that are resilient to a changing environment.
Plant scientist Zach Lippman – a CSHL PI profile
July 16, 2019
Lippman’s research focuses on the process of flowering and flower production in plants, major contributors to reproductive success and crop yield.
An essay from the President: Biology for the planet
May 16, 2019
CSHL plant scientists are looking for solutions to the biggest questions in agriculture as environments are reshaped by climate change.
Profile: Doreen Ware champions the plant genome
May 8, 2019
Molecular biologist Doreen Ware uses computer science to parse out the genetic roadmaps of plants.
Cryptic mutation is cautionary tale for crop gene editing
May 6, 2019
Unexpected interactions between mutations can be a thorn in the side for plant breeders. Scientists unveil what drove one infamous “cryptic” mutation.
How do plants sense their environment?
April 30, 2019
CSHL plant scientist Ullas Pedmale details how plants sense the world we share.
Cold Spring Harbor Laboratory announces exclusive license with plant breeding start-up Inari
April 16, 2019
CSHL announced a licensing agreement with partner Inari, a company that is advancing plant breeding by tapping nature’s genetic diversity.
Rob Martienssen wins Martin Gibbs Medal for plant research
April 15, 2019
CSHL Professor Rob Martienssen wins the 2019 Martin Gibbs Medal for his contributions to plant biology.
To protect stem cells, plants have diverse genetic backup plans
April 15, 2019
Experts discover how an essential genetic circuit found in all flowering plants, regardless of species, is protected in startlingly different ways.
Crop yield in maize influenced by unexpected gene ‘moonlighting’
April 1, 2019
Yield of the maize plant is tied to activity of a gene called RAMOSA3, but new evidence suggests the gene performs other unexpected functions
The year of CRISPR
December 26, 2018
A look at the various labs across CSHL that utilize CRISPR in their research, and the groundbreaking discoveries they help uncover.
New regulators of nitrogen use in plants identified
October 24, 2018
A team of CSHL plant biologists identify gene regulators that may help plants utilize nitrogen better and prevent excess nitrogen in soil.
CRISPR could bring groundcherries to market
October 1, 2018
CSHL Professor Zachary Lippman uses CRISPR to make the groundcherry more suitable for large-scale farming
Groundwork for playing with the architecture of plants
August 24, 2018
Scientists discover how to stimulate stem cells in a specific spot to aid in future agriculture.
Base Pairs Episode 16.5: Fuels of the fuels
August 16, 2018
Rob Martienssen explains how genetic modification and advances in technology factor into the future of fuel as well as cinematic sci-fi fuels.
Base Pairs Episode 16: Big plans for a tiny plant
July 16, 2018
As temperatures around the globe continue to rise, scientists are working hard to develop solutions that deal with the consequences of climate change.
Big plans for a tiny plant
July 15, 2018
On this episode of Base Pairs, Rob Martienssen discusses how duckweed could be the next biofuel and help combat climate change
Energy from thin air: Basic research to biofuels
June 12, 2018
A plant scientist and an industry innovator discuss how simple duckweed may help address climate change and energy needs.
Prof. Zachary Lippman named Blavatnik Award finalist
May 30, 2018
Professor Zachary Lippman was chosen as a Finalist in Life Sciences for the 2018 Blavatnik National Awards
CSHL’s Zachary Lippman named HHMI Investigator
May 23, 2018
CSHL Professor Zach Lippman is selected as an HHMI investigator, in recognition of his innovative work in the field of plant genetics
How do plants know when to flower?
May 18, 2018
Colorful flowers are a sure sign of spring. But how do flowering plants know that it is time to bloom?
One experiment: Twice the tomatoes
March 22, 2018
Zachary Lippman’s team found a way to boost tomato yield by digging into the plant’s genome and fine-tuning its branching patterns.
The secret to tripling the number of grains in sorghum and perhaps other staple crops
February 26, 2018
A simple genetic modification can triple grain production in sorghum, a drought-tolerant plant that is an important source of food and animal feed.
Counting chromosomes: Plant scientists solve a century-old mystery about reproduction
January 18, 2018
Geneticists have solved a century-old mystery by discovering a remarkable mechanism that enables plants to count their chromosomes
Light, cryptochromes, action…
November 21, 2017
Assistant Professor Ullas Pedmale has received the National Institutes of Health "Outstanding Investigator Award."
The changing relationship between humans and plants, it’s complicated
November 3, 2017
CSHL scientists Dave Jackson, Doreen Ware, and Zachary Lippman give their perspectives on the rapidly changing relationship between humans and plants.
New application of CRISPR breaks age-old yield barriers in crops
September 15, 2017
CSHL's Lippman lab has harnessed the untapped power of genome editing to improve agriculture.
Plant geneticists develop a new application of CRISPR to break yield barriers in crops
September 14, 2017
Mutating regulatory regions varies yield traits the way a dimmer switch controls a light bulb
Demystifying GMOs
July 21, 2017
We've all heard of GMOs, but what exactly ARE they? Hannes, a plant scientist from CSHL's Dave Jackson lab, explains genetic modification.
Base Pairs Episode 10.5: Tomato baby and its family
July 14, 2017
Plant scientist Zachary Lippman tells stories from the field of bizarre tomatoes, intensely hot peppers, and giant pumpkins.
Tomato baby and its family
July 14, 2017
This episode of Base Pairs talks to Associate Professor Zachary Lippman about all the weird mutations that naturally occur in plants.
CRISPR vs. climate change
June 15, 2017
This episode of Base Pairs looks at the issues climate change may bring to agriculture and how CRISPR can help overcome them.
Detailed new ‘reference’ genome for maize shows the plant has deep resources for continued adaptation
June 12, 2017
A new, much more detailed reference genome for maize, or corn, as it is called in the U.S., provides new insight into the crop.
Fine-tuning mutant genes unleashes long-trapped potential in tomato
May 19, 2017
A team of plant geneticists at CSHL publish a paper in Cell demonstrating how bringing together beneficial traits can have negative consequences.
Fine-tuning dosage of mutant genes unleashes long-trapped yield potential in tomato plants
May 18, 2017
Understanding gene interactions can enable breeders to break existing productivity barriers in agriculture.

David Jackson
My lab studies genes and signals in cells that regulate the growth and shape of plants. We have discovered several genes that control plant architecture by exerting an influence on stem cells. By identifying the genes that control the number of stem cells in corn plants, for example, we’ve discovered a means of boosting the yield of that vital staple.

Zachary Lippman
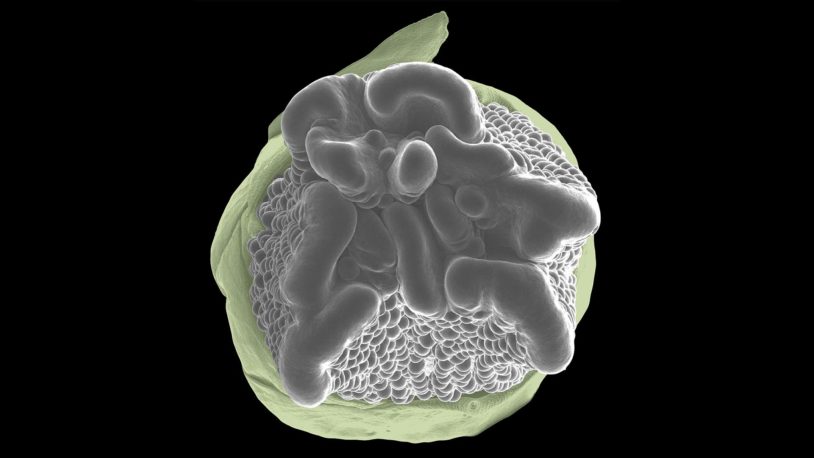
My research team studies when and where, and how many branches, flowers, and fruits are produced on plants. All of plant development depends on small groups of stem cells at the tips of shoots known as meristems. By studying the genes that control stem cell production and maturation over space and time, within and between different developmental contexts, we are able to manipulate plant architecture and reproduction to improve crop yields.

Rob Martienssen
Chromosomes are covered with chemical modifications that help control gene expression. I study this secondary genetic code - the epigenome - and how it is guided by small mobile RNAs in plants and fission yeast. Our discoveries impact plant breeding and human health, and we use this and other genomic information to improve aquatic plants as a source of bioenergy.

W. Richard McCombie
Over the last two decades, revolutionary improvements in DNA sequencing technology have made it faster, more accurate, and much cheaper. We are now able to sequence up to 10 trillion DNA letters in just one month. I harness these technological advancements to assemble genomes for a variety of organisms and probe the genetic basis of neurological disorders, including autism and schizophrenia, better understand cancer progression and understand the complex structures of the genomes of higher plants.

Ullas Pedmale
Unlike animals, plants neither have specific organs that see or hear various stimuli, yet, plants are sensitive to their surrounding environment and modify their development according to various external signals. My lab studies how the environment of a plant modulates its growth and development. Understanding environmental control of growth will have far-reaching implications for agriculture, energy production, and many other human activities.

Doreen Ware
When we think of evolution, we often think about physical changes, like a plant developing broader leaves to collect more solar energy. Such evolution actually occurs within the plant’s DNA. I am using computational analysis and modeling to visualize how plant genomes have evolved over time, particularly those of staple crops. We are learning from this work to improve the range and yield of modern plants.