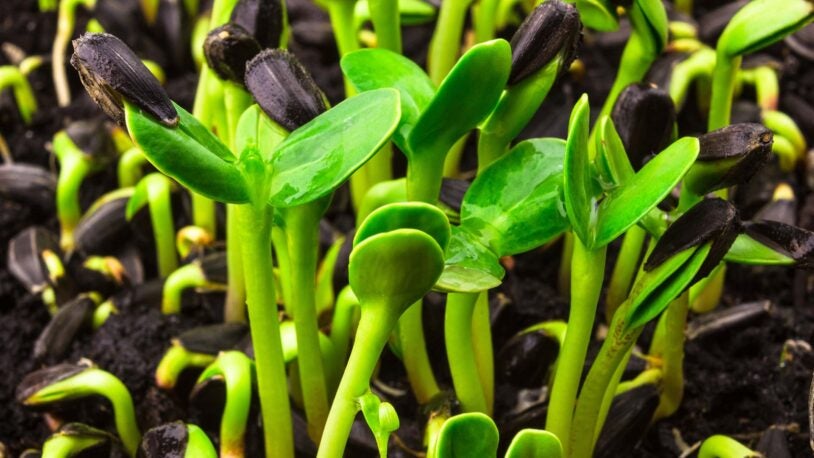
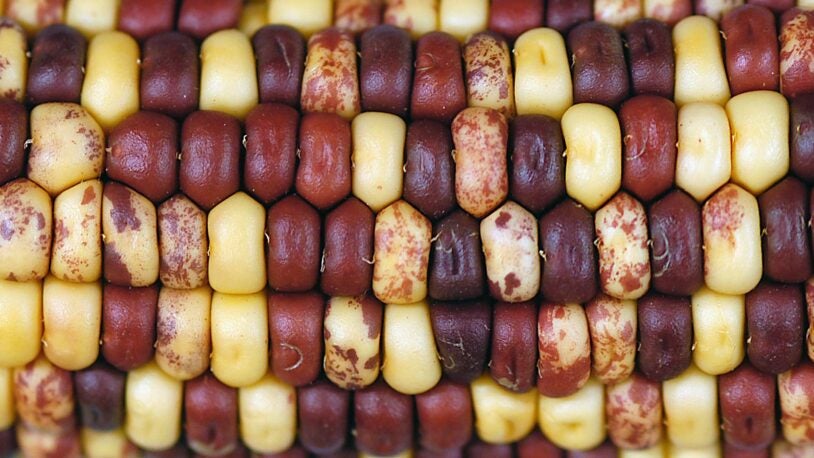
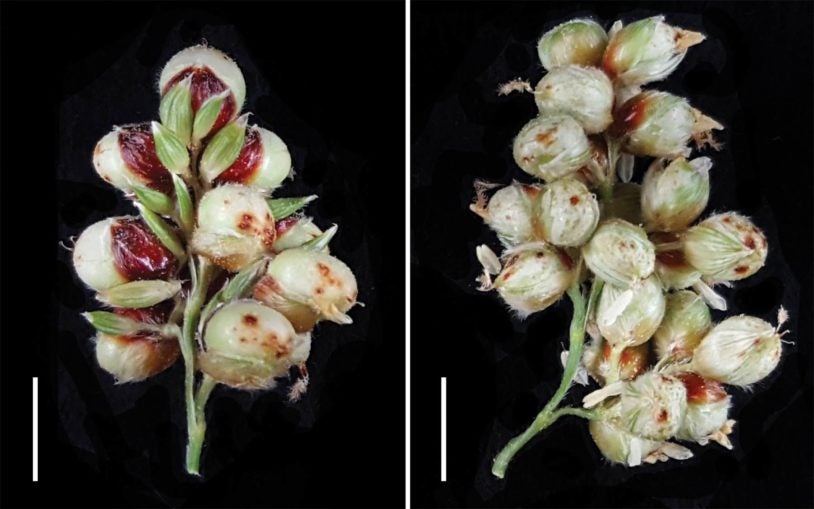
When we think of evolution, we often think about physical changes, like a plant developing broader leaves to collect more solar energy. Such evolution actually occurs within the plant’s DNA. I am using computational analysis and modeling to visualize how plant genomes have evolved over time, particularly those of staple crops. We are learning from this work to improve the range and yield of modern plants.
Using multidisciplinary approaches that combine computational analysis, modeling, and prediction with experimental verification, Doreen Ware’s lab seeks a deeper understanding of the evolution of genome sequences in plants and their implications for agricultural improvement. By looking comparatively across the genomes of plants in the same lineage, they seek answers to the following questions: How are genes conserved and lost over time? What are the fates of duplicated genes? What is the impact of structural variation on phenotypic variation? Ware’s team also studies gene regulation in plants, focusing on gene regulatory networks, targeting transcription factors and microRNA genes with the objective of understanding how these parts of the plant genome work together in determining spatial and temporal expression of genes. The lab had an important role in the project to produce a haplotype map reference genome of maize, spearheading the most comprehensive analysis of the crop yet. This has provided important information on the variation of the reference genome, as well as comparative data showing changes in the genome acquired through domestication and breeding. They have devoted special attention to examining diversity within maize, grape, and tomato, aiming to accelerate the development of strategies to introduce new germplasm that is needed to meet demands of increasing population and a changing environment. The lab also has brought fully sequenced genomes into an integrated data framework, to enhance the power of their comparative studies. This past year, Ware was named as its principal investigator for the National Science Foundation-funded Gramene project, a comparative genomics resource for agriculturally important crops and models to support sustainable food and fuel production. Ware, as principal investigator for plants, has also helped lead an effort funded by the Department of Energy to create—out of many separate streams of biological information—a single, integrated cyber-“knowledgebase” for plants and microbial life.
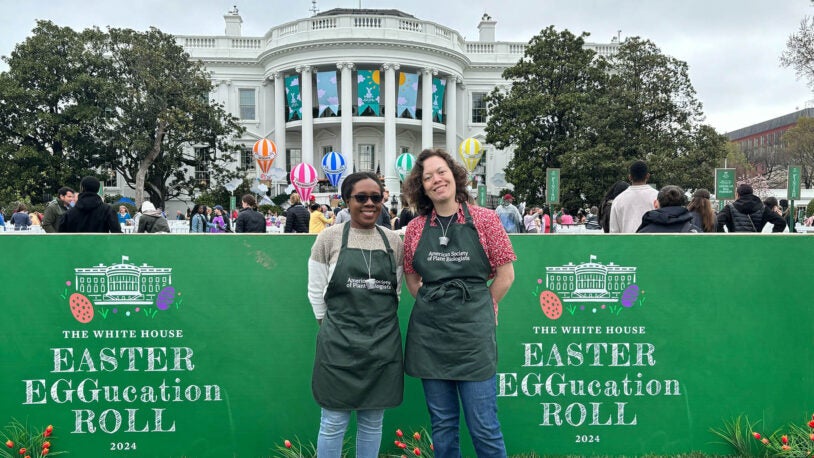
CSHL goes to the White House for Easter EGGucation
April 8, 2024
CSHL plant scientists taught kids the basics of plant biology and its role in the environment at an event hosted by First Lady Jill Biden.
New CSHL website brings together sorghum researchers
May 9, 2022
Plant researchers and breeders are now using a website created by CSHL to get the latest intel on sorghum crop research.
The race to protect sweet corn
April 22, 2022
Breeding a variety that can withstand disease and taste better too
Do you have the dirt on plant research?
March 31, 2022
New research is constantly sprouting. Take this quiz and test your plant knowledge.
The secret history of corn is revealed in its genome
August 5, 2021
For the first time, scientists have assembled in-depth maps of dozens of corn genomes, filling in gaps related to key agricultural traits.
Teachers make genomes more useful… from home
May 21, 2020
Researchers and educators participated in an online annotation jamboree to help researchers understand the corn genome.
CSHL investigators rank among world’s most highly cited
December 11, 2019
Seven researchers affiliated with CSHL are among the scientists producing the top 1 percent of the most highly-cited research in the world.
Researchers double sorghum grain number to improve food supply
October 30, 2019
A set of hormone-controlling genes may be the key to doubling grain number in sorghum plants.
Leading discovery
May 20, 2019
Current discoveries about DNA and human genome position CSHL scientists to make life-changing breakthroughs that will improve the human condition.
An essay from the President: Biology for the planet
May 16, 2019
CSHL plant scientists are looking for solutions to the biggest questions in agriculture as environments are reshaped by climate change.
All Publications
Time-series multi-omics analysis of micronutrient stress in Sorghum bicolor reveals iron and zinc crosstalk and regulatory network conservation
22 May 2025 | Plant Biology
Mishra, A; Bhat, A; Kumari, S; Sharma, R; Braynen, J; Tadesse, D; El Alaoui, S; Seaver, S; Grosjean, N; Ware, D; Xie, M; Paape, T;
GrameneOryza: a comprehensive resource for Oryza genomes, genetic variation, and functional data
4 Apr 2025 | Database : the journal of biological databases and curation | 2025
Wei, Sharon; Chougule, Kapeel; Olson, Andrew; Lu, Zhenyuan; Tello-Ruiz, Marcela; Kumar, Vivek; Kumari, Sunita; Zhang, Lifang; Olson, Audra; Kim, Catherine; Gladman, Nick; Ware, Doreen;
Genome sequence assembly and annotation of MATA and MATB strains of Yarrowia lipolytica
25 Mar 2025 | bioRxiv
Zali, Narges; El Damerash, Osama; Chougule, Kapeel; Lu, Zhenyuan; Ware, Doreen; Stillman, Bruce;
A pan-gene catalogue of Asian cultivated rice
20 Feb 2025 | bioRxiv
Contreras-Moreira, Bruno; Sharma, Eshan; Saraf, Shradha; Naamati, Guy; Gupta, Parul; Elser, Justin; Chebotarov, Dmytro; Chougule, Kapeel; Lu, Zhenyuan; Wei, Sharon; Olson, Andrew; Tsang, Ian; Lodha, Disha; Zhou, Yong; Yu, Zhichao; Zhao, Wen; Zhang, Jianwei; Amberkar, Sandeep; Sue-Ob, Kawinnat; Martin, Maria; McNally, Kenneth; Ware, Doreen; Sun, Zhi; Deutsch, Eric; Copetti, Dario; Wing, Rod; Jaiswal, Pankaj; Dyer, Sarah; Jones, Andrew;
MaizeCODE reveals bi-directionally expressed enhancers that harbor molecular signatures of maize domestication
30 Dec 2024 | Nature Communications | 15(1):10854
Cahn, Jonathan; Regulski, Michael; Lynn, Jason; Ernst, Evan; de Santis Alves, Cristiane; Ramakrishnan, Srividya; Chougule, Kapeel; Wei, Sharon; Lu, Zhenyuan; Xu, Xiaosa; Ramu, Umamaheswari; Drenkow, Jorg; Kramer, Melissa; Seetharam, Arun; Hufford, Matthew; McCombie, W; Ware, Doreen; Jackson, David; Schatz, Michael; Gingeras, Thomas; Martienssen, Robert;
Building a FAIR data ecosystem for incorporating single-cell transcriptomics data into agricultural genome to phenome research.
28 Nov 2024 | Frontiers in Genetics | 15:1460351
Kapoor, Muskan; Ventura, Enrique; Walsh, Amy; Sokolov, Alexey; George, Nancy; Kumari, Sunita; Provart, Nicholas; Cole, Benjamin; Libault, Marc; Tickle, Timothy; Warren, Wesley; Koltes, James; Papatheodorou, Irene; Ware, Doreen; Harrison, Peter; Elsik, Christine; Yordanova, Galabina; Burdett, Tony; Tuggle, Christopher;
Gapless assembly of complete human and plant chromosomes using only nanopore sequencing
6 Nov 2024 | Genome Research
Koren, Sergey; Bao, Zhigui; Guarracino, Andrea; Ou, Shujun; Goodwin, Sara; Jenike, Katharine; Lucas, Julian; McNulty, Brandy; Park, Jimin; Rautiainen, Mikko; Rhie, Arang; Roelofs, Dick; Schneiders, Harrie; Vrijenhoek, Ilse; Nijbroek, Koen; Nordesjo, Olle; Nurk, Sergey; Vella, Mike; Lawrence, Katherine; Ware, Doreen; Schatz, Michael; Garrison, Erik; Huang, Sanwen; McCombie, William; Miga, Karen; Wittenberg, Alexander; Phillippy, Adam;
Functional diversification within the heme-binding split-barrel family
10 Oct 2024 | Journal of Biological Chemistry | :107888
Grosjean, Nicolas; Zhang, Lifang; Kumaran, Desigan; Xie, Meng; Fahey, Audrey; Santiago, Kassandra; Hu, Fangle; Regulski, Michael; Blaby, Ian; Ware, Doreen; Blaby-Haas, Crysten;
GRAS Family Transcription Factor Binding Behaviors in Sorghum bicolor, Oyrza, and Maize
25 Sep 2024 | bioRxiv
Gladman, Nicholas; Kumari, Sunita; Fahey, Audrey; Regulski, Michael; Ware, Doreen;
High-quality chromosome scale genome assemblies of two important Sorghum inbred lines, Tx2783 and RTx436
Sep 2024 | NAR Genomics and Bioinformatics | 6(3):lqae097
Wang, Bo; Chougule, Kapeel; Jiao, Yinping; Olson, Andrew; Kumar, Vivek; Gladman, Nicholas; Huang, Jian; Llaca, Victor; Fengler, Kevin; Wei, Xuehong; Wang, Liya; Wang, Xiaofei; Regulski, Michael; Drenkow, Jorg; Gingeras, Thomas; Hayes, Chad; Armstrong, J; Huang, Yinghua; Xin, Zhanguo; Ware, Doreen;