Understanding gene interactions can enable breeders to break existing productivity barriers in agriculture
Cold Spring Harbor, NY — Breeding in plants and animals typically involves straightforward addition. As beneficial new traits are discovered—like resistance to drought or larger fruits—they are added to existing prized varieties, delivered via cross-breeding. Yet every once in a while, adding a beneficial new trait can result in a net subtraction, due to processes deeply hidden within the interactions of genes underlying existing and newly added traits.
Interaction among genes—both positive and negative—is called epistasis (“ep-i-STAY-sis”). Today, a team of plant geneticists at Cold Spring Harbor Laboratory (CSHL) publishes a paper in Cell demonstrating how bringing together beneficial traits can have negative consequences. They discover and dissect a case of negative epistasis in a variety of the tomato plant. But they also show how to exploit this knowledge to derive untapped yield potential from the plant. They do so by cross-breeding specimens of the plant carrying different “dosages” of gene variants responsible for positive and negative traits.
“Our study provides the first example of which we’re aware of a domestication gene that hindered crop improvement,” says CSHL Associate Professor Zachary Lippman, who led the research. “This work illustrates how gene dosage can be exploited to fine-tune and improve major yield traits. It shows that by identifying and dissecting similar cases of negative epistasis in plant and animal breeding, we may be able to break existing productivity barriers in agriculture.”
A tale of two mutations
Nearly all tomatoes grown today—possibly everyone you’ve ever eaten—carries a gene mutation that likely arose 8,000 to 10,000 years ago, at the dawn of agriculture. This ancient mutation caused early forms of domesticated tomato plants to produce fruits with larger green leafy caps on top. We’ll never know for sure why ancient tomato growers liked their tomatoes with larger caps, but we do know that they selected for such plants, and crossed them so that the trait became part of the genome in most modern tomato varieties.
The second mutation tracked down by Lippman’s team that interacts with this ancient mutation was discovered in the mid-twentieth century, first spotted in domesticated tomatoes planted in a field owned by the Campbell Soup Company. This second mutation caused two dramatic changes. Called jointless, the newly mutated gene changed the elbow-like bend in the stem leading to the plant’s flowers—a natural joint called an abscission zone. These mutant tomato plants, as “jointless” implies, had no joints.
This was a wonderful trait for industrial growers, who by the postwar period wanted to harvest fruit with mechanical pickers. It’s much easier for a harvester (mechanical or human) to cleanly detach the tomato from the vine in the jointless variety, since the elbow in the stem vanishes and the green cap on top of the fruit remains on the plant. When tomatoes are tossed into a container as they are picked, those with caps and bits of residual stem often puncture other tomatoes nearby. So growers liked jointless and breeders sought to exploit it immediately.
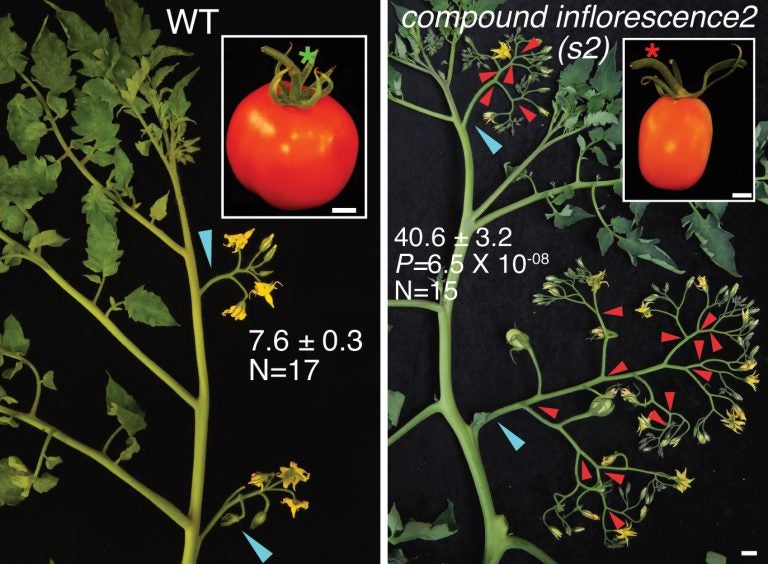
But the ancient mutation that enlarged the green caps prevented the easy adoption of jointless. For reasons unknown to breeders and growers, and not known until the newly published research from CSHL, jointless plants frequently had too many of the branches that make flowers, known as inflorescences.
“You would think—correctly—that more inflorescence branches mean more flowers,” says Lippman, “and in fact that’s one way to get more yield. But only up to a point. If tomato or any other plant makes too many flowers, the plant doesn’t have enough resources to turn all those flowers into fruits. The result is actually a decline in fertility.”
Commercial breeders eventually found a way to counteract the unwanted branching, which enabled them to take advantage of the mutation that generated jointless stems. The result was inflorescences shaped in a way familiar to anyone who has ever seen tomatoes on the vine. The fruits hang off single stems, arrayed in a linear zig-zag pattern.
Reversing a negative gene interaction
That is exactly what he and colleagues succeeded in doing. Their key discovery: realizing that the ancient mutation that makes larger green caps in most modern tomato plants interacts with the jointless mutation first noticed in the Campbell Soup field some decades ago. This interaction is a telling example of “negative epistasis,” says Lippman—“a case of two mutations, one ancient and one recent, both considered beneficial when they were selected, when placed in the same plant have a negative effect.”
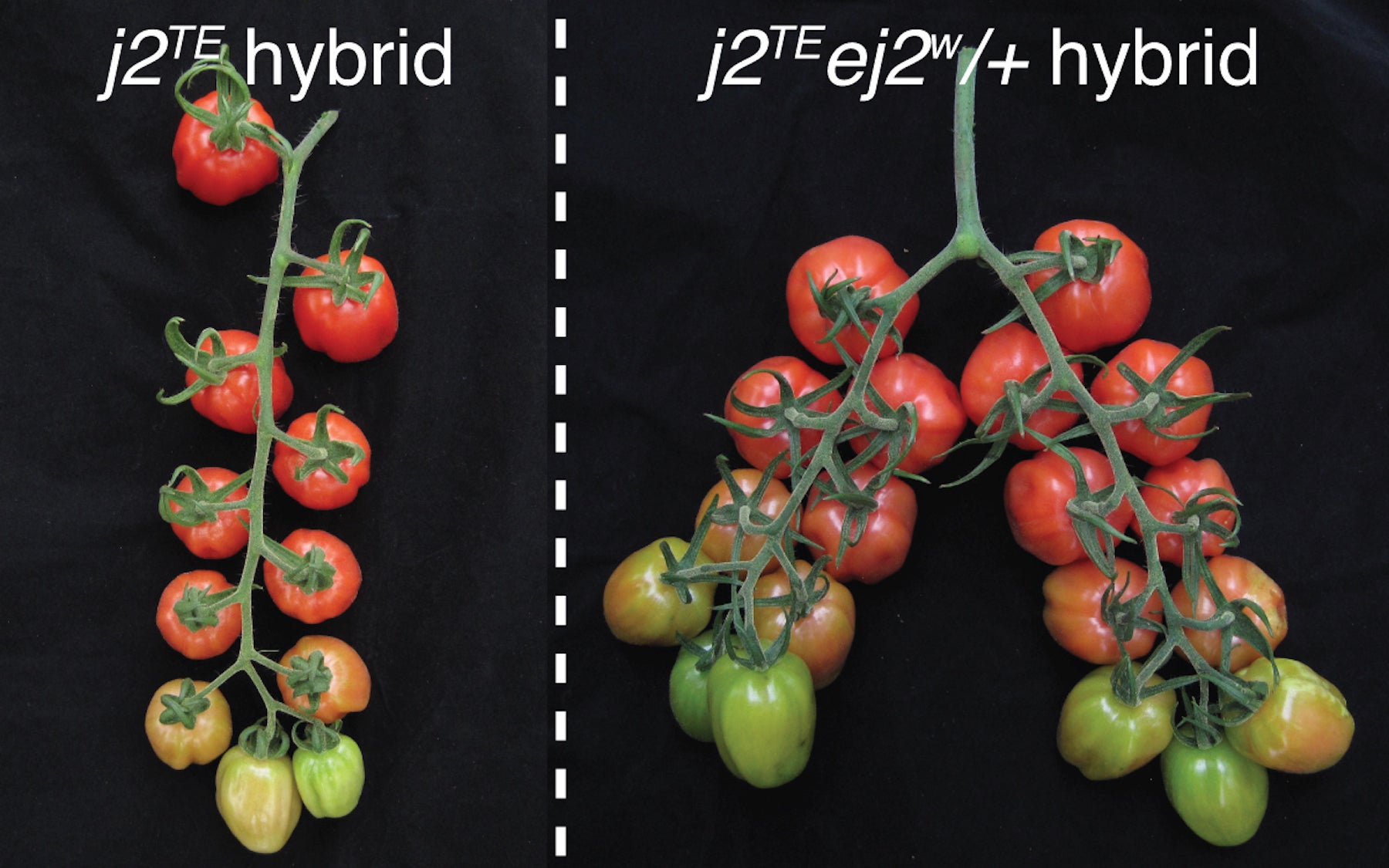
By totally negating the trait conferred by jointless that was unwanted—that explosion of branching—“we think breeders missed some important potential in yield.” The team was able to capture and exploit this potential by adjusting the “dosage” of the gene responsible for the size of the green cap. Rather than zeroing-out its impact on branching, like commercial growers did, they were able to make genetic crosses with variants carrying different versions (called alleles) of the gene, so that only a fraction of its power to spur branching is expressed.
The result is a weakly branching tomato variety that is also jointless, hence easy to harvest. Yet they also have more fruits, simply because they develop just a few more branches and flowers.
This strategy of identifying, neutralizing and potentially exploiting negative epistasis, says Lippman, could help improve other crops, and also perhaps domesticated animals in which there may be more hidden gene interactions with less obvious negative effects that are preventing breeders and farmers from realizing their full genetic potential.
Written by: Peter Tarr, Senior Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
The research discussed here was supported by: an EMBO Long-Term Fellowship (ALTF 1589-2014); a National Science Foundation Postdoctoral Research Fellowship in Biology Grant (IOS-1523423); the Next-Generation BioGreen 21 Program (SSAC, PJ01188301); the National Science Foundation Plant Genome Research Program (IOS-1564366); a European Research Council – Advanced Grant entitled YIELD (ERC-294691); the US-Israel Binational Agricultural Research & Development fund; and the National Science Foundation Plant Genome Research Program (IOS-1237880).
Citation
“Bypassing negative epistasis on yield in tomato imposed by a domestication gene” appears online in Cell May 18, 2017. The authors are: Sebastian Soyk, Zachary H. Lemmon, Matan Oved, Josef Fisher, Katie L. Liberatore, Soon Ju Park, Anna Goren, Ke Jiang, Alexis Ramos, Esther van der Knaap, Joyce Van Eck, Dani Zamir, Yuval Eshed, and Zachary B. Lippman. The paper can be accessed at: http://www.cell.com/cell/home
Principal Investigator

Zachary Lippman
Professor & HHMI Investigator
Jacob Goldfield Professor of Genetics
Director of Graduate Studies
Ph.D., Watson School of Biological Sciences at Cold Spring Harbor Laboratory, 2004

2 thoughts on “Fine-tuning dosage of mutant genes unleashes long-trapped yield potential in tomato plants”
Comments are closed.