Leemor Joshua-Tor
Professor, Director of Research & HHMI Investigator
W.M. Keck Professor of Structural Biology
Cancer Center Member
Ph.D., The Weizmann Institute of Science, 1991
[email protected] | 516-367-8821
Our cells depend on thousands of proteins and nucleic acids that function as tiny machines: molecules that build, fold, cut, destroy, and transport all of the molecules essential for life. My group is discovering how these molecular machines work, looking at interactions between individual atoms to understand how they activate gene expression, DNA replication, and small RNA biology.
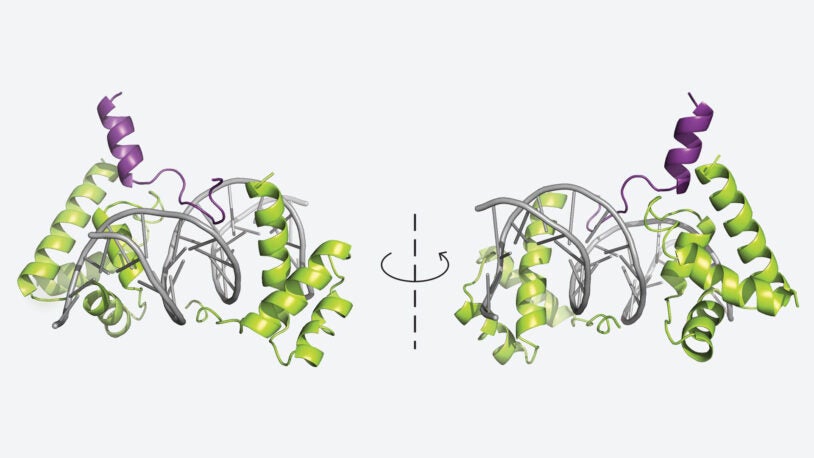
In Leemor Joshua-Tor’s lab, researchers study the molecular basis of nucleic acid regulatory processes using the tools of structural biology and biochemistry. One such regulatory process is RNA interference (RNAi), in which a small double-stranded RNA triggers gene silencing. Joshua-Tor and her team offered critical insight when they solved the crystal structure of the Argonaute protein and identified it as the long-sought Slicer. They then went on to explore the mechanism of the slicing event. The structure of human Argonaute 2 (hAgo2) bound to a microRNA (miRNA) guide allowed Joshua-Tor and her colleagues to understand how mRNA is cleaved during RNAi. This year, members of the Joshua-Tor lab explored the function of a very similar protein, called Argonaute 1, that has no slicing ability, even though it is almost identical in structure to the slicing hAgo2. Using biochemical methods and mutational analysis, they were able to identify key parts of the protein that are required for slicing activity. The lab also studies the generation of PIWI-interacting RNAs (piRNAs), which serve to protect the genome of germ cells. With colleagues in the Hannon lab, Joshua-Tor’s team also determined the structure and function of Zucchini, a key nuclease in the initial generation of piRNAs in fruit flies. In other work, the lab is exploring the mechanisms of heterochromatin formation and gene silencing through the study of a protein complex called RNA-induced initiation of transcriptional gene silencing (RITS). Joshua-Tor is also well known for her work on the E1 helicase enzyme, which acts to unwind DNA strands during the DNA replication process.
Dr. Leemor Joshua-Tor honored with Mildred Cohn Award from ASBMB
Member of the National Academy of Sciences
Member of the American Academy of Arts and Sciences
2018 Mildred Cohn Award in Biological Chemistry, ASBMB
2014 ACE Women’s Network, New York, Women in Science and Education Leadership Award
Fellow of the American Association for the Advancement of Science (AAAS)
2007 Dorothy Crowfoot Hodgkin Award (Inaugural award), The Protein Society
1996 Beckman Young Investigator Award
Shapeshifting cancers’ masters, unmasked
November 24, 2025
New research from CSHL’s Vakoc lab helps explain the bizarre behavior of certain pancreatic cancers and lung cancers that can change identities.
Joshua-Tor joins American Academy of Sciences and Letters
November 13, 2025
The prestigious group counts among its ranks Sir Salman Rushdie and three Nobel laureates, all with a background in chemistry.
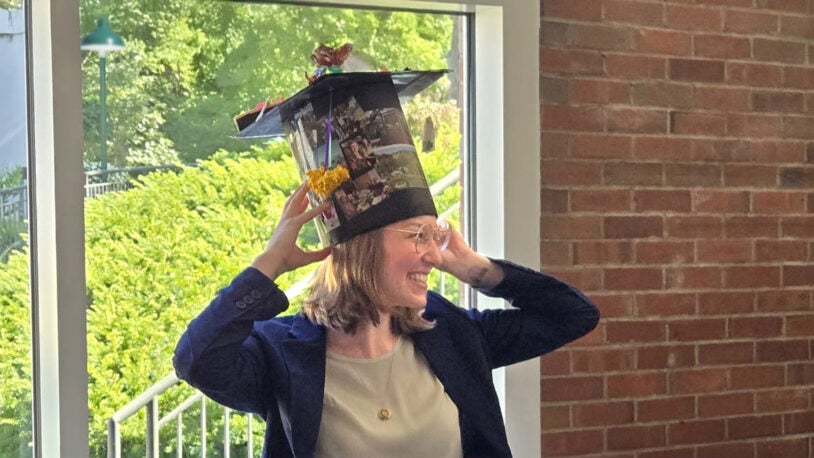
A graduation cap like no other
September 10, 2025
Group arts and crafts projects provide the Joshua-Tor lab a fun and memorable way to recognize its graduate students.
A recipe to reverse cancer’s sweet tooth
June 16, 2025
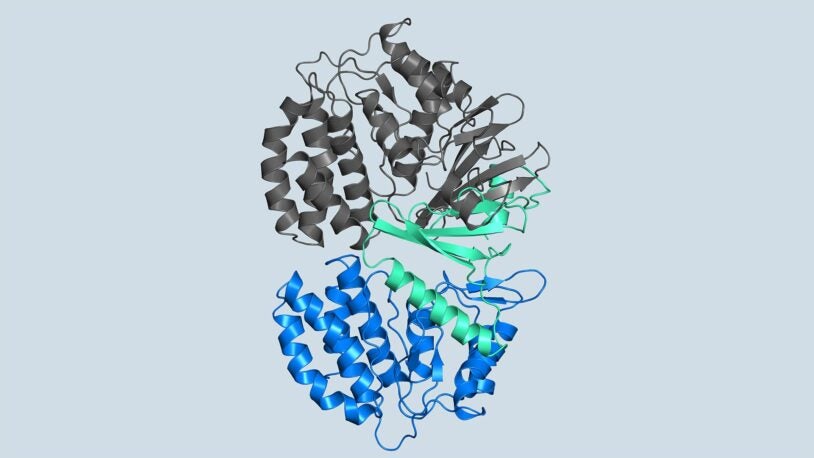
Researchers reveal the first 3D structures of the kinase FN3K in different states, which may inform future therapeutics for a variety of cancers.
At the Lab: The human Microprocessor
May 27, 2025
There’s a Microprocessor inside you. Actually, there are trillions. Only they’re not computer processors. They’re much smaller and far more complex.
The Microprocessor inside you
December 2, 2024
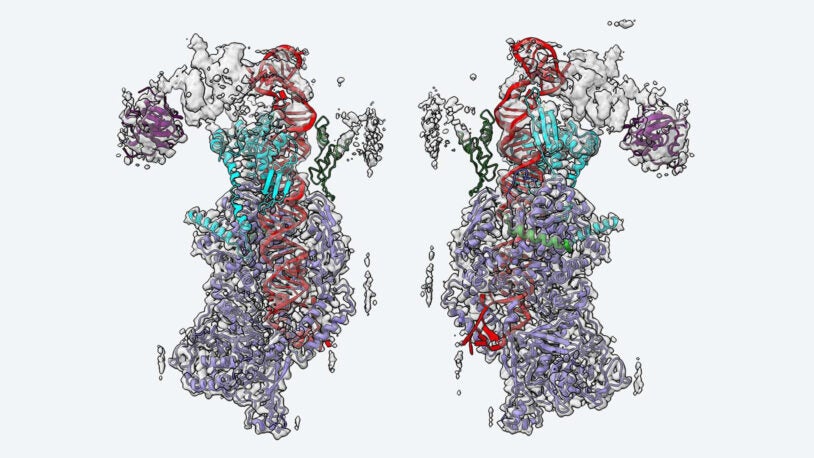
Stunning new images from CSHL’s Joshua-Tor lab reveal how the protein complex interacts with differently shaped primary microRNAs.
The 2024 CSHL Raft Race
November 12, 2024
Ten teams brave cool harbor waters, the hot August sun, and new boat-building rules in the 9th annual CSHL Raft Race.
RNA splicing’s spotters
June 10, 2024
RNA therapeutics pioneer CSHL Professor Adrian Krainer has discovered a link between two important regulator proteins, SRSF1 and DDX23.
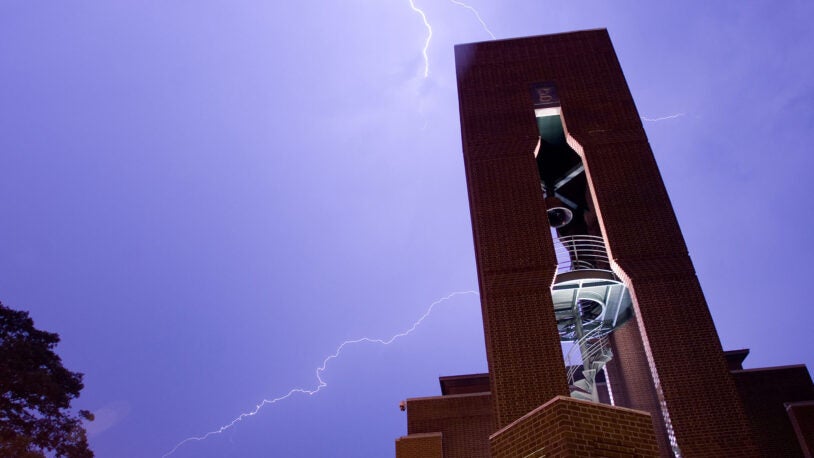
Hazen Tower
April 25, 2024
The Italian-style bell tower anchors CSHL’s Neuroscience Center. Its bell has rung out every hour on the hour, from 8 a.m. to 8 p.m., since 1991.
Up close and personal with cryo-EM
April 11, 2024
CSHL’s Cryo-Electron Microscopy course teaches the next generation of scientists to study life at the atomic level.
Selected Publications
The structural landscape of Microprocessor-mediated processing of pri-let-7 miRNAs
1 Oct 2024 | Molecular Cell | :S1097-2765(24)00741
Garg, Ankur; Shang, Renfu; Cvetanovic, Todor; Lai, Eric; Joshua-Tor, Leemor;
Chromatin remodeling of histone H3 variants by DDM1 underlies epigenetic inheritance of DNA methylation
24 Aug 2023 | Cell | 186(19):4100-4116.e15
Lee, Seung; Adams, Dexter; Ipsaro, Jonathan; Cahn, Jonathan; Lynn, Jason; Kim, Hyun-Soo; Berube, Benjamin; Major, Viktoria; Calarco, Joseph; LeBlanc, Chantal; Bhattacharjee, Sonali; Ramu, Umamaheswari; Grimanelli, Daniel; Jacob, Yannick; Voigt, Philipp; Joshua-Tor, Leemor; Martienssen, Robert;
A shape-shifting nuclease unravels structured RNA
23 Feb 2023 | Nature Structural & Molecular Biology
Meze, Katarina; Axhemi, Armend; Thomas, Dennis; Doymaz, Ahmet; Joshua-Tor, Leemor;
Target binding triggers hierarchical phosphorylation of human Argonaute-2 to promote target release
31 May 2022 | eLife | 11
Bibel, Brianna; Elkayam, Elad; Silletti, Steve; Komives, Elizabeth; Joshua-Tor, Leemor;
All Publications
Repeated emergence of giant microRNA hairpins across invertebrates
9 Sep 2025 | Cell Reports | 44(9):116243
Javad-Zada, Mir-Mammad; Zou, Xiaodong; Okamura, Katsutomo; Shang, Renfu; Garg, Ankur; Lee, Ethan; Joshua-Tor, Leemor; Fromm, Bastian; Lai, Eric;
Structural and mechanistic insights into Dis3L2–mediated degradation of structured RNA
2 Aug 2025 | bioRxiv
Matos, Rute; Garg, Ankur; Costa, Susana; Pereira, Patrícia; Arraiano, Cecília; Joshua-Tor, Leemor; Viegas, Sandra;
The molecular basis of Human FN3K mediated phosphorylation of glycated substrates
22 Jan 2025 | Nature Communications | 16(1):941
Garg, Ankur; On, Kin; Xiao, Yang; Elkayam, Elad; Cifani, Paolo; David, Yael; Joshua-Tor, Leemor;
The molecular basis of Human FN3K mediated phosphorylation of glycated substrate
5 Aug 2024 | bioRxiv
Garg, Ankur; On, Kin; Xiao, Yang; Elkayam, Elad; Cifani, Paolo; David, Yael; Joshua-Tor, Leemor;
SRSF1 interactome determined by proximity labeling reveals direct interaction with spliceosomal RNA helicase DDX23 (correction)
2024 | Proceedings of the National Academy of Sciences of the United States of America | 121(30)
Segovia, Danilo; Adams, Dexter; Hoffman, Nickolas; Tepes, Polona; Wee, Tse-Luen; Cifani, Paolo; Joshua-Tor, Leemor; Krainer, Adrian;
SRSF1 interactome determined by proximity labeling reveals direct interaction with spliceosomal RNA helicase DDX23
21 May 2024 | Proceedings of the National Academy of Sciences of the United States of America | 121(21):e2322974121
Segovia, Danilo; Adams, Dexter; Hoffman, Nickolas; Safaric Tepes, Polona; Wee, Tse-Luen; Cifani, Paolo; Joshua-Tor, Leemor; Krainer, Adrian;