The birth and evolution of open access data banks”
Before the age of modern cloud computing and the Internet, collecting and distributing scientific data was a challenging task. Researchers from different countries had their own ways of cataloging their findings. Computers were rare, expensive, and inaccessible.
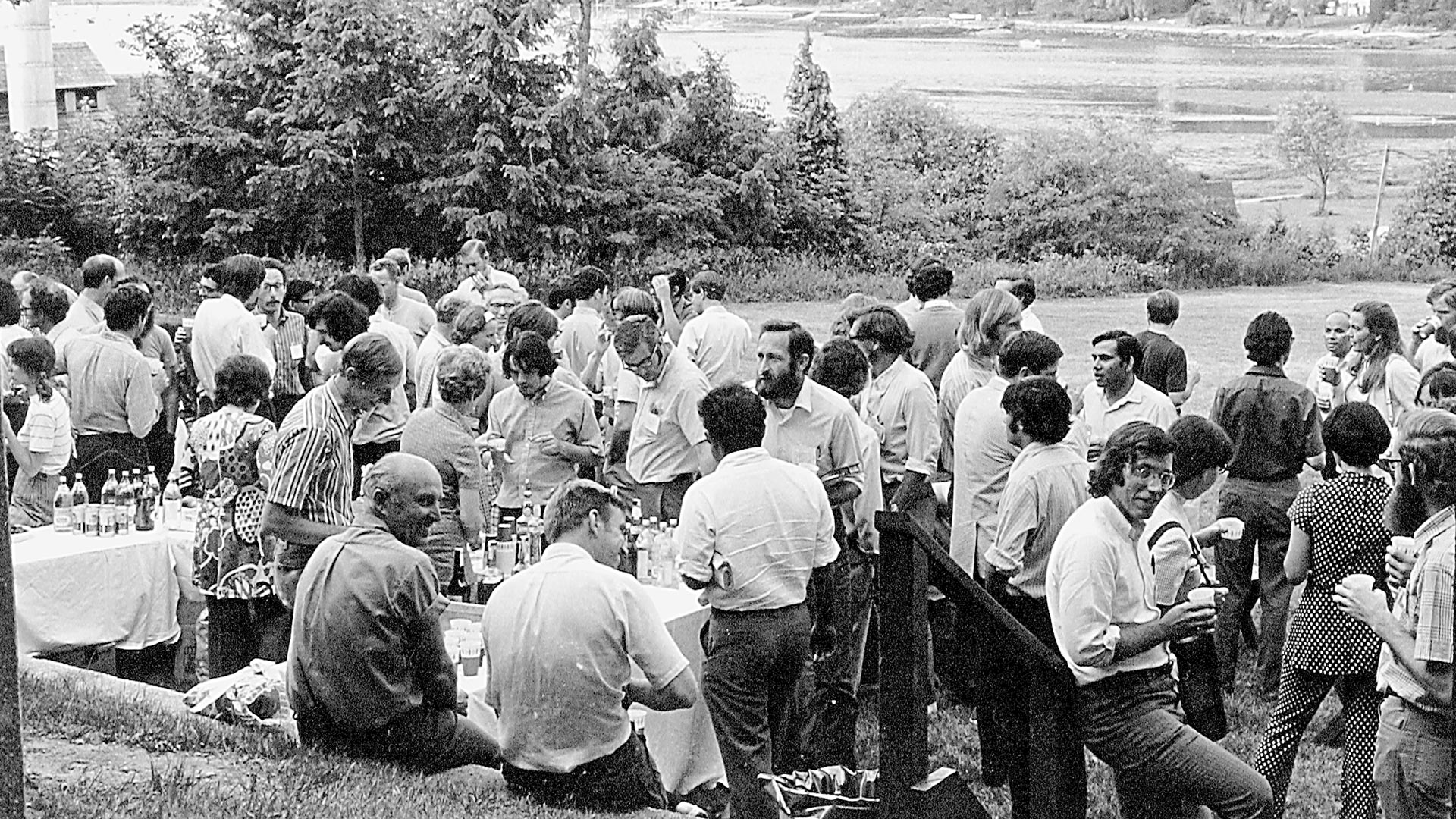
The Cold Spring Harbor Laboratory (CSHL) June 1971 symposium, titled “Structure and Function of Proteins at the Three-Dimensional Level,” brought together protein scientists, including four Nobel laureates. Max Perutz, who won the 1962 Nobel Prize for discovering the structure of hemoglobin, sparked an informal discussion on how to collect and distribute protein structure data. Symposium attendees agreed on the need for an open access data storage system. Walter Hamilton, a chemist developing graphic technologies and remote computing at the nearby Brookhaven National Laboratory, volunteered to establish a digital database, a first-of-its-kind public protein data bank.
The first barrier—speaking the same language
A protein’s function depends on its ability to coil, kink, or bend into a specific three-dimensional shape. Structural biologists need to know the precise location of each atom. In 1971, scientists from different places used different coordinate systems, which made it hard to share their findings. Agreeing to speak only one language—one standard coordinate system—would allow people from all around the world to understand each other’s work.
Though only a few protein structures were solved at the time, scientists dreamed of cataloging dozens, hundreds, perhaps even thousands of structures. With the help of several leading structural biologists, Hamilton wrote software to store atomic coordinate files into a common format.
One language, many databases
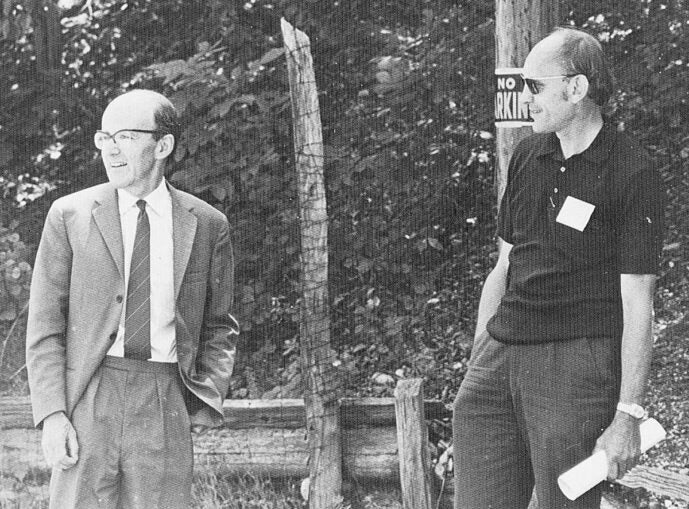

Hamilton’s database—the Protein Data Bank (PDB)—was officially launched in October 1971. It started with less than a dozen structures and was the first open-access digital data resource in biology or medicine. The PDB began receiving support from the National Science Foundation in 1975, while other data banks were set up in Europe and Japan. By 1998, major journals (Nature, Science, and PNAS) required structures to be deposited to a publicly accessible data bank as a condition of publication. That same year, the Research Collaboratory for Structural Biology (RCSB) took over the data bank management. Five years later, a new international foundation was formed to combine protein data banks and manage the PDB, called the Worldwide PDB (wwPDB). The wwPDB ensures that “the PDB is freely and publicly available to the global community.”
The data bank grew exponentially since its start, with a collection of 100 structures in 1982, 1,000 in 1993, 10,000 in 1999, and 100,000 in 2014. By the end of 2021, the PDB held over 185,000 structures.
Building in transparency
The PDB is a model for global cooperation and transparency, allowing structural biologists to see what others are studying and lend a helping hand. Once researchers outside the structural biology field started to see these benefits, they formed their own data banks, some of which started at meetings or in labs at CSHL. Here are four examples of open access data banks, which serve as foundational resources in their fields.
The Human Genome Project
The Human Genome Project (HGP) was one of the first large-scale international biology projects. Scientists hammered out the path for the HGP at a 1989 Banbury meeting. Launched in October 1990, the goal of the collaborative project was to sequence the three billion letters of the human genome. The Genome Database, created by Johns Hopkins University with funding from the Howard Hughes Medical Institute, played an important role in organizing genomic data from a consortium of researchers from 20 institutes, including CSHL.
ENCODE
After the Human Genome Project was completed in 2003, the National Human Genome Research Institute shifted its attention to the Encyclopedia of DNA Elements (ENCODE). ENCODE is a comprehensive catalog of functional elements in the human and mouse genomes. By uncovering DNA segments that affect specific functions and identifying them across all individuals, researchers can standardize the genome and pinpoint where variations and diseases may arise from person to person. CSHL Professor Thomas Gingeras, along with an international consortium of approximately 500 scientists, reported the completion of ENCODE Phase 3 in 2020.
Plant genomes
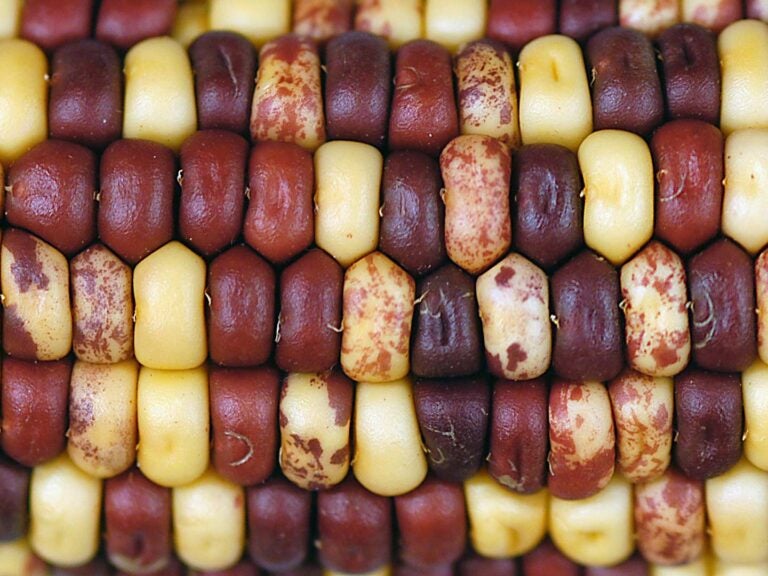
CSHL played a key role in the early stages of sequencing the genomes of plants. In 1994, CSHL Professor and HHMI Investigator Rob Martienssen, along with colleagues Mike Bevan and Joe Ecker, organized a Banbury Center meeting (pdf) to persuade the National Science Foundation to fund the sequencing of the model plant, Arabidopsis. The initiative was successful, and many online collections of plant genomes began to crop up. CSHL contributed to many subsequent efforts. For example, CSHL Adjunct Professor Doreen Ware and her colleagues, including Martienssen and CSHL Professor W. Richard McCombie, first mapped the corn genome in 2009. In 2021, they published more data, filling in the gaps in the previous map.
Brain maps
Neuroscientists are also reaping the benefits of online data banks for sharing information on brain cells. In 2013, the U.S. government invested $100 million into the Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative, which aims to understand human brain anatomy. The initiative sought to take a census of every single cell in the brain through the BRAIN Initiative Cell Census Network. The network’s extensive open access resources are managed by the Allen Institute.
Databases to the rescue
The PDB is now a critical piece of infrastructure for the international biology research community. To celebrate the 50th anniversary of its founding, scientists around the world held conferences and virtual symposia. CSHL researchers like Professor and HHMI Investigator Leemor Joshua-Tor, Professor Hiro Furukawa, and President and CEO Bruce Stillman study protein structures in new and evolving ways; they and other contributors continue to grow the PDB. The CSHL Meetings & Courses Program is training new generations of structural biologists. Open access data banks are considered foundational elements to any large scientific enterprise.

Transparent and accessible data served us well during the COVID-19 pandemic. Even before January 2020, when the World Health Organization declared the COVID-19 outbreak a “Public Health Emergency of International Concern,” scientists were at work sharing information. Scientific journals usually take weeks, months, or even years to process raw manuscripts into published peer-reviewed articles. However, the CSHL preprint servers bioRxiv and medRxiv provided open access platforms to post unpublished research manuscripts within hours or days. The preprint servers became the COVID-19 research communications hub for scientists, journalists, and the general public. Thanks to that near-instant exchange of findings, the coronavirus was isolated, sequenced, and classified in weeks; vaccines were quickly developed.
Open access data banks give communities of scientists the tools to study proteins, analyze genomes, map brains, and fight coronaviruses. The troves of data they are assembling today will prepare the world for the health emergencies of tomorrow.
Written by: Jasmine Lee, Content Developer/Communicator | publicaffairs@cshl.edu | 516-367-8845