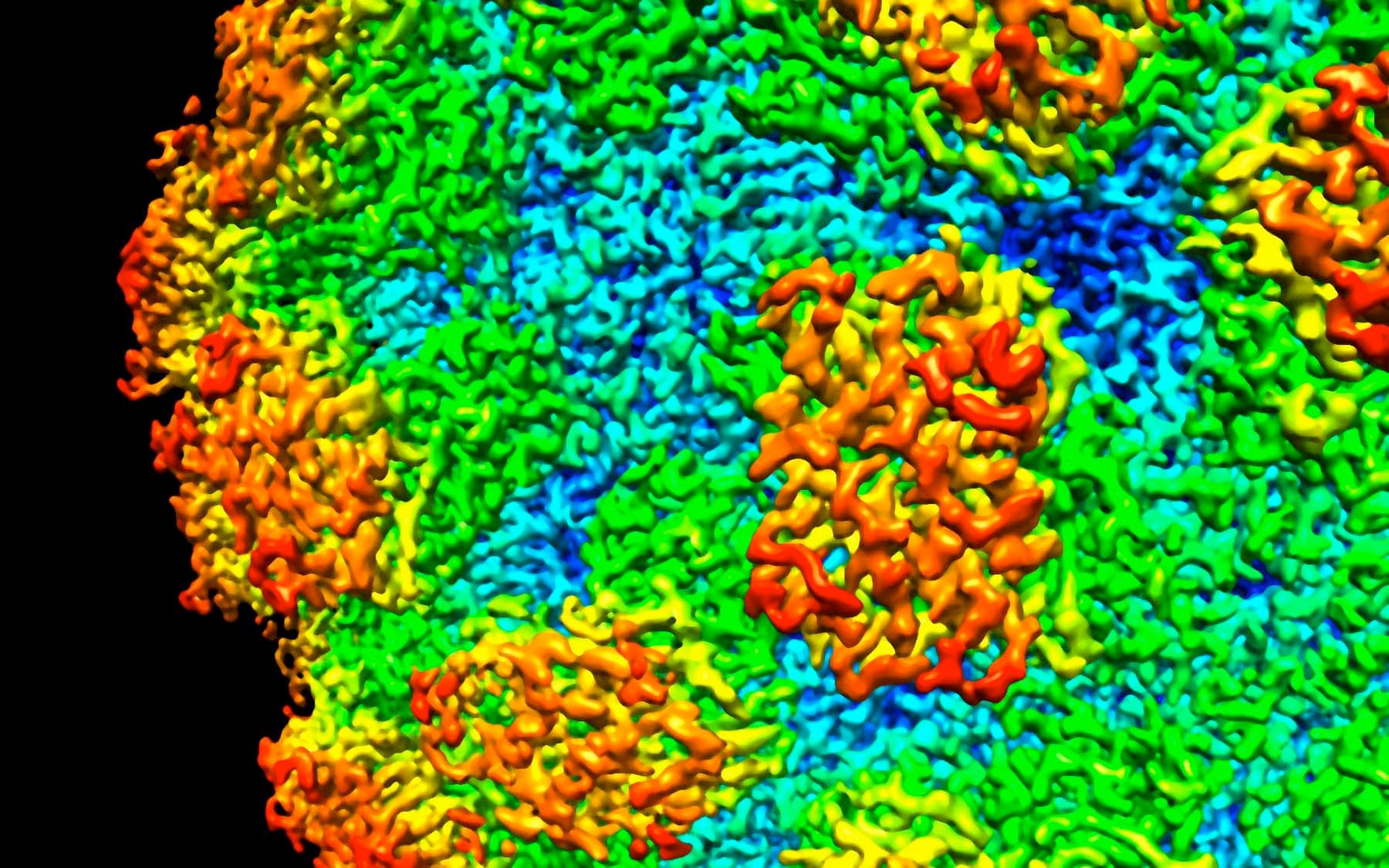
Viruses encapsulate their genetic cargo in a shell of proteins. These shells can form in various shapes and sizes.
The shapes of the virus shells are important for developing vaccines against future viral infections. They provide target outlines of the virus for the immune system, which then creates a type of protein called antibodies that can bind to the viruses’ shells so they can no longer interact with human cells.
Noroviruses account for 58 percent of all stomach bug outbreaks in the United States and cause 685 million cases worldwide each year. There are currently no antiviral treatments or vaccines against it.
New research from Professor Leemor Joshua-Tor’s lab at Cold Spring Harbor Laboratory, published in PNAS, used a cryo-electron microscope to solve the shapes and sizes of various strains of noroviruses.
Joshua-Tor’s lab mainly studies the functions and structural biology of proteins and nucleic acids. The noroviruses provided a nice entry into hi-resolution cryo-electron microscopy.
“There has been a structure of just one strain of norovirus called GI.1 that was solved by X-ray crystallography,” said James Jung, a postdoctoral fellow in Joshua-Tor’s lab and the first author on the study. “Other than that, we didn’t know if other strains would assemble in a similar manner.”
With the cryo-electron microscope, the team was able to show broad structural differences between four strains of noroviruses from two broad genotypes: the most prevalent GII.4, GII.2, GI.7, and GI.1.
“You can raise the antibodies from more than one virus form or have a mixture, but you have to be aware that there is such variability,” said Joshua-Tor. Knowledge of the different forms of norovirus shell structures can help better inform future vaccine developments and antiviral treatments.
Written by: Charlotte Hu, Content Developer/Communicator | publicaffairs@cshl.edu | 516-367-8455
About

Leemor Joshua-Tor
Professor, Director of Research & HHMI Investigator
W.M. Keck Professor of Structural Biology
Cancer Center Member
Ph.D., The Weizmann Institute of Science, 1991