In food-poisoning outbreaks, like those caused by noroviruses in hospitals, schools, restaurants and on cruise ships, infections occur and spread quickly. Also known as “stomach flu” and “winter vomiting bug,” noroviruses are the most common and extremely contagious breed of stomach bugs, accounting for 58% of all outbreaks in the United States and causing 685 million cases worldwide each year. Norovirus illnesses are estimated to rack up $60 billion in costs for care and lost productivity globally. There are currently no approved therapeutics for treating these infections.
A virus is a different brand of villain than cancer or even bacteria. Viruses aren’t technically alive because they can’t grow and need a host cell to replicate. Their genetic information is simply protected by a sheath of proteins, otherwise known as shells, or capsids. Elements on the norovirus shells that jut out, called spikes, are what allows the virus to attach to receptors on human host cells. The viruses then hijack the human cell and use it as a reproduction factory, multiplying exponentially.
Developing vaccines against viruses involves creating specific antibodies that will recognize the norovirus shells and bind to them so they can no longer interact with human cells.
“In order to develop vaccines and anti-virals more effectively, we need to understand what the capsid shapes actually look like, and if there are any differences between different strains,” said James Jung, a postdoctoral fellow in Professor and HHMI Investigator Leemor Joshua-Tor’s lab at Cold Spring Harbor Laboratory (CSHL).
So far, the assembled virus shell structures have only been available for a single minor strain of norovirus, named GI.1, imaged using X-ray crystallography. To broaden the knowledge and understanding of norovirus architecture, Jung, Joshua-Tor, and a team of biologists sought to solve the high-resolution structures of four different outbreak strains of noroviruses with a powerful cryo-electron microscope that uses electrons as the light source. “The resolution was so good that we could actually see water molecules bound as well,” Jung said. This allowed them to make out the intricate architecture of virus shells in near-atomic detail. Their findings were published in the journal PNAS.
Introduction to the workings and possibilities of the cryo-electron microscope (credit: Dennis Thomas)
Making of a norovirus
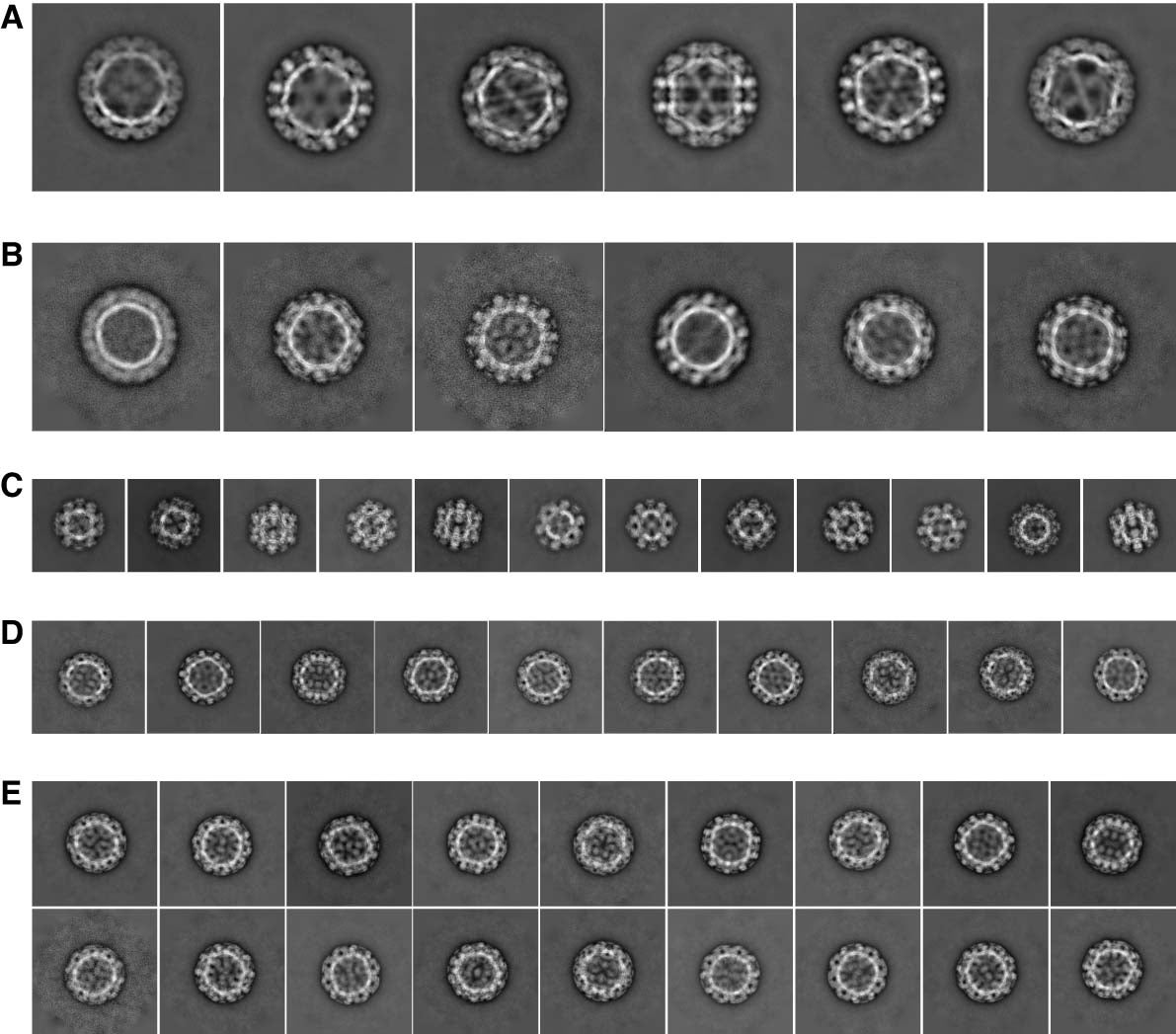
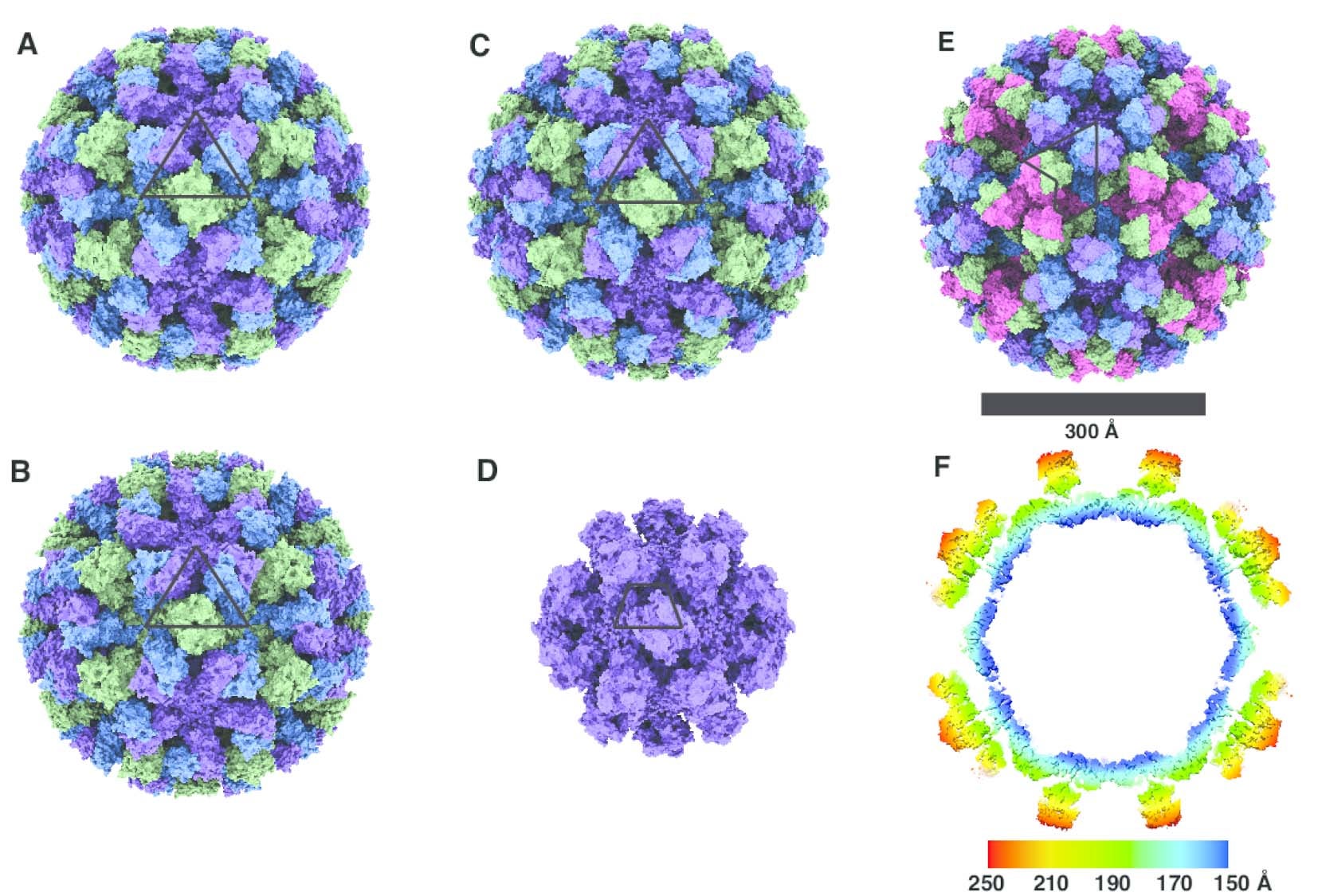
Using cryo-EM, Jung was able to show broad structural differences between four strains of noroviruses from two broad genotypes: the most prevalent GII.4, GII.2, GI.7, and GI.1. He prepared the virus shells frozen in a glass-like state that preserved the shells in their native-like forms and protected them from the bombarding electrons by sealing them in a vacuum. He then recorded them in different orientations in 2D movies to get sharper, motion-stabilized views. By sorting the shells in different views and computationally putting the 2D projections together, he was able to reconstruct virus shells into 3D structures.
The final product almost resembles the Hoberman sphere toy synonymous with childhood: colorful, skeletal, and collapsible balls that were inspired by physics.

By solving the three-dimensional architecture of the virus shells, Jung gleaned new insights that could help guide development of much-needed therapeutics to fight norovirus infection. “Previously, it was thought that the norovirus shells exist in a single-sized assemblies consisting of 180 building blocks and 90 surface spikes. What we found was an unexpected mixture of different shell sizes and shapes. We found a smaller form, which consists of just 60 building blocks with 30 surface spikes placed further apart. We also found larger shells made out of 240 building blocks with 120 surface spikes that are lifted significantly above the base of the shell and form a two-layered architecture that would interact differently with the human cells,” he said.
The spikes on the shell are the parts that interact with the host. Jung found that the spacing and orientation of the spikes varied significantly across the different strains of noroviruses. “That means it’s going to have different interactions with the human cells,” Jung explained. “It also means that the way the antibody recognizes and binds the virus shells is going to be different. This is significant because when it comes to formulating the vaccines, to ensure the maximum efficacy, they have to be formulated in such a way that takes into account all of these variations across strains and structural forms.”
The zinc link
The hi-resolution imaging also allowed the team to look at “the very tip top of the spike that wasn’t observed at all in an older study,” said Joshua-Tor. “That’s the part that at least we know binds to one coreceptor on human cells.”
From what the team observed, there were two flexible protein loops on the spike of the GII.2 shell that are likely antigenic or capable of binding to an antibody. While the specific protein receptor that human noroviruses bind to is yet to be defined, it is known that to attach to human cells, the virus needs to bind another molecule as a coreceptor. This molecule is the human histo-blood group antigens (HBGA), which determine the A/B/O blood group. In the absence of HBGA, viruses were unable to attach to human cells and replicate.
“Right next to the site that’s involved in histo-blood antigen binding was a zinc metal ion,” Jung said. It seems to him that the metal ion is acting like a linchpin, holding the two loops together in the correct conformation so the virus can properly latch onto the human cell. When this metal ion was removed using a zinc-chelating agent, three-quarters of the shells were destroyed.
Professor Joshua-Tor further elaborated that this means if scientists want to keep the virus shells stable and raise antibodies that can block the histo-blood group antigen binding site, it may be important to add metal to the vaccine formulation.
Currently, one vaccine candidate for norovirus treatment is in phase two of clinical trials. Jung and Joshua-Tor hope that by drawing up a blueprint of norovirus shell designs, they can help better inform future vaccine developments that take into account the variations across strains.
Written by: Charlotte Hu, Content Developer/Communicator | publicaffairs@cshl.edu | 516-367-8455
About

Leemor Joshua-Tor
Professor, Director of Research & HHMI Investigator
W.M. Keck Professor of Structural Biology
Cancer Center Member
Ph.D., The Weizmann Institute of Science, 1991