Cold Spring Harbor, NY — A doctor treating a patient with a potentially fatal metastatic breast tumor would be very pleased to find, after administering a round of treatment, that the primary tumor had undergone a change in character—from aggressive to static, and no longer shedding cells that can colonize distant organs of the body. Indeed, most patients with breast and other forms of cancer who succumb to the illness do so because of the cancer’s unstoppable spread.
A team at Cold Spring Harbor Laboratory (CSHL) today publishes preclinical data based on experiments conducted in mice suggesting the promise of a novel drug directed against a novel target in malignant mammary tumors. Although the work is at the first stage of development, it excites team leader David L. Spector, Ph.D., because in mouse models of human metastatic breast cancer it has produced “differentiation of an aggressive primary tumor, redirecting it from a highly proliferative state, and reducing its metastatic activity by 70 percent.”
Spector, Professor and Director of Research at CSHL, is using an unusual kind of drug to hit an unusual kind of cancer target. Most anti-cancer drugs are chemicals or proteins that target proteins in cells. In this case, both the target and the candidate drug are nucleic acids.
The target is called Malat1. It is a member of an understudied class of nucleic acids that biologists call long non-coding RNAs (lncRNAs). These RNA molecules are copied from portions of the genome that do not “code” for proteins. Malat1’s function in cells has not yet been determined. But for over a decade it has been known to have unusual qualities, compared with other long non-coding RNAs localized in the cell nucleus.
In addition to its especially long length, Malat1 was found to be among the most abundant lncRNAs. As Spector observes, “it takes a great deal of energy for cells to make large quantities of this RNA, so we tend to assume it is doing something important.” The argument is augmented by a series of findings demonstrating that Malat1 is even more abundant than usual in some classes of malignant tumor cells.
In an earlier study, Spector’s team knocked out the Malat1 gene in mice and curiously, these mice had no apparent abnormalities, a fact that would seem improbable if Malat1 did, in fact, have an important function. Spector, however, was intrigued by earlier reports of the upregulation of the RNA in certain cancers. Therefore, his team, led by postdoctoral fellow Gayatri Arun, set out to discover what would happen if mice that model human metastatic breast cancer were bred with the mice lacking Malat1.
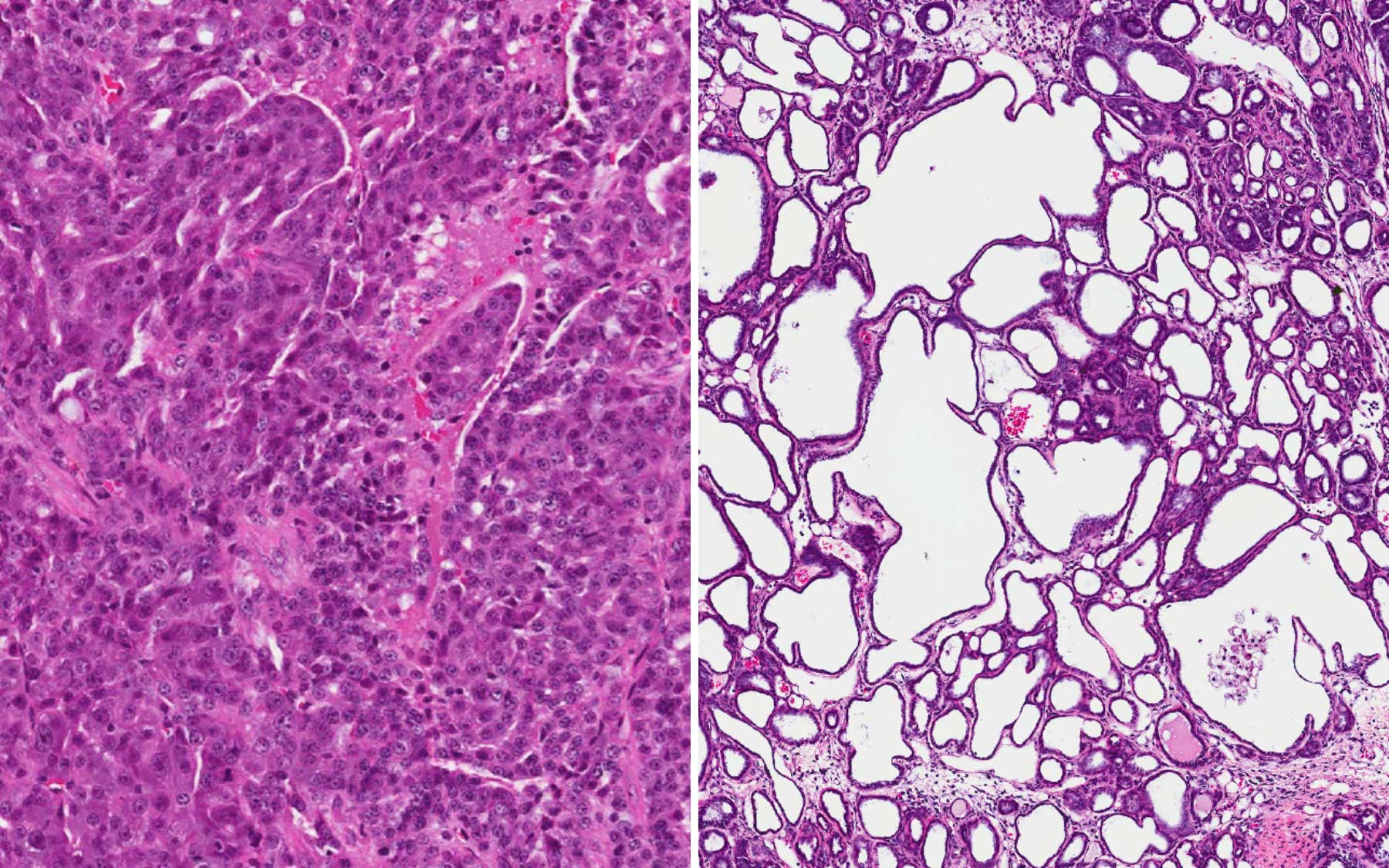
“We got an amazing result,” Spector says. “By removing Malat1—this one, single long non-coding RNA—we made a dramatic impact on the primary breast tumors in these mice. The tumors took on a wholly new character.” Aggressive breast cancer tumors differentiated and took on cyst-like characteristics. They filled with fluid, which, the team noted, contained various proteins including milk proteins, demonstrating that some of the tumor cells had differentiated. Just as important, in these mice lacking Malat1, there was nearly no metastasis.
Gene editing experiments like this—in which a gene is knocked out—are not performed in humans. The question Spector and colleagues considered was how to design a drug that would mimic the effects of knocking out the Malat1 gene. Perhaps, they reasoned, such a drug might produce some of the remarkable therapeutic effects seen in the knock-out mice.
In collaboration with Ionis Pharmaceuticals (Carlsbad, CA) they utilized an ASO—an antisense oligonucleotide—a nucleic acid designed by virtue of its complementary nucleotide sequence to bind to the Malat1 lncRNA at a precise location. The ASO-Malat1 hybrid stimulates a naturally occurring cellular enzyme that degrades the Malat1 lncRNA.
The ASO was administered to a line of mice that mimicked human metastatic breast cancer. Although these mice had an active Malat1 gene, its RNA was degraded in animals that were treated with the Malat1 ASO. As in the knock-out mice, aggressive tumors failed to thrive and were reduced to low-grade tumors looking more like cysts; metastasis, while not completely abolished as in the knock-out mice, was reduced some 70 percent.
“We are very pleased and excited by this result,” Spector says, “because it suggests that these metastatic tumors have a dependency on Malat1—they can’t thrive without it. And very importantly, targeting Malat1 should not have a deleterious effect on any normal cells.” At least in this mouse model, Spector adds, “normal, non-cancerous cells don’t seem to care a bit if they have Malat1 or not. Only the cancer cells seem to require it.”
The team conducted additional experiments in which mouse breast tumors were sampled to generate so-called organoids—spheres of cells that grow in three-dimensional culture. The organoids were used to further explore Malat1’s role in breast cancer cells. These experiments, led by postdoctoral fellows Gayatri Arun and Sarah Diermeier, indicated Malat1 “is touching multiple nodes—gene programs that determine what cells do.” In aggressive cancer cells with Malat1 knocked out completely, or degraded by the ASO treatment, the organoids are no longer highly active; they stop moving and pack tightly together—suggesting, perhaps, how targeting Malat1 reduces metastatic potential.
The team is continuing with their experiments, to study the activity of this and other therapeutic ASOs in human breast cancer, with the hope of later moving the research into human clinical trials.
Written by: Peter Tarr, Senior Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
The research described here was supported by National Cancer Institute grant 5P01CA013106-Project 3, the Babylon Breast Cancer Coalition, and a Cancer Center Support Grant to Cold Spring Harbor Laboratory (2P30CA45508).
Citation
“Differentiation of Mammary Tumors and Reduction in Metastasis Upon Malat1 LncRNA Loss” appears online in Genes & Development on December 23, 2015. The authors are: Gayatri Arun, Sarah Diermeier, Martin Akerman, Kung-Chi Chang, J. Erby Wilkinson, Stephen Hearn, Youngsoo Kim, A. Robert MacLeod, Adrian R. Krainer, Larry Norton, Edi Brogi, Mikala Egeblad and David L Spector. The paper can be obtained at: http://genesdev.cshlp.org/
Principal Investigator

David L. Spector
Professor
Robert B. Gardner Jr., Professor
Cancer Center Member
Ph.D., Rutgers University, 1980