Sometimes science can lead to unexpected discoveries in unexpected places. That’s what Cold Spring Harbor Laboratory (CSHL) Professor David Spector experienced this year. For nearly four decades, Spector has been researching cancer—the United States’ second-leading cause of death. But this year, he found himself studying the first—heart disease.
Spector’s research at CSHL focuses on how genes are expressed, or turned on and off. He also studies a type of molecule called long non-coding RNAs (lncRNAs).

RNA is a nucleic acid present in all living cells. Like DNA, it’s made up of compounds called nucleotides. Many types of RNA molecules help build proteins, but lncRNAs don’t serve this function. Instead, they have varying jobs, including chaperoning proteins to genes to turn them on or off. At 200 nucleotides or longer, lncRNAs are thought of as cell regulators.
Spector’s lab studies lncRNAs often expressed in breast cancer. He also researches lncRNAs during differentiation—the process through which stem cells turn into more specialized cells.
So this year, when two separate projects on gene expression led Spector’s lab into cardiac research, he was understandably surprised.
“My lab is absolutely not known for studying cardiac differentiation or development,” he says. “We’ve been interested both in cancer and in the process of differentiation. Of course, there are parallels between those two processes.”
Both heart studies were published within the last two months, and both identify how the expression of certain genes can lead to cardiac problems in mice. Not just a common killer in the US, heart disease is one of the leading causes of death worldwide. Understanding how the heart develops and the different genetic causes of heart-related conditions could one day lead to new treatments for these often-fatal diseases.
It looks like a heart attack. The young animal should not die at 10 to 12 weeks of age.”
CSHL Postdoc Rasmani Hazra
The first project builds upon previous research from Spector’s lab, which focused on lncRNAs that are highly expressed in mouse embryonic stem cells (mESCs). A few years ago, the team identified 30 of these RNAs called Platrs (short for pluripotency-associated transcripts). This study, published in Developmental Cell, honed in on one of those—Platr4.
Spector’s lab set its sights on Platr4 because humans have several possible counterparts in our cells. “That made it particularly interesting,” he explains.
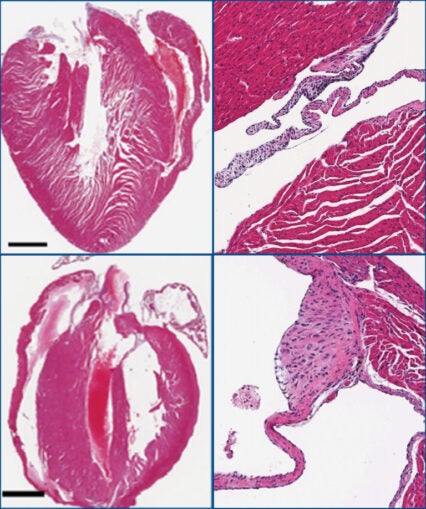
Spector says he didn’t know what Platr4’s exact function was before this study. He assumed it worked directly in stem cells, helping to keep them actively dividing. Instead, the team discovered that Platr4 helps stem cells form into heart cells. It acts as a chaperoning lncRNA, bringing molecules to a gene called Ctgf and turning on that gene. Ctgf is crucial for the development of heart valves—and without Platr4, Ctgf doesn’t function.
“We had no inkling that the result would be what we found. The science led us there,” Spector says. “The findings were a big surprise.”
CSHL postdoctoral fellow Rasmani Hazra spearheaded the study in Spector’s lab. She first suspected Platr4 was involved in heart development when she began removing it from mESCs to see how they were affected. When Hazra switched off Platr4 in mouse embryos and let them develop, a significant number of the mice died from heart valve problems. After extracting and examining the hearts, Hazra found defects that restricted their ability to pump blood.
A control group of mice also expressed Platr4 at the embryo stage, but didn’t have it switched off and grew up with perfectly healthy hearts. After these animals were born, Platr4 turned off naturally with no ill effects. Interestingly, in both groups of mice, Platr4 was only present in developing embryos, yet its removal at this early stage had disastrous results for the living organisms’ beating hearts.
“It looks like a heart attack,” Hazra says. “The young animal should not die at 10 to 12 weeks of age.”
Humans have three possible Platr4 equivalents in our cells, all located on chromosome 4. Further research is needed to explore these lncRNAs’ roles in human heart valve development and function. But clinical case studies have already linked the deletion of chromosome 4 to cardiac and structural defects in humans.
Spector and Hazra suspect a defective Platr4 counterpart may act as a biological marker for valvular heart disease. If they’re correct, it could lead to earlier diagnoses.
Valvular heart disease affects 2.5% of the US population and claims the lives of some 25,000 people in the country each year. It can be successfully treated with medicines like beta-blockers, but quick detection is critical. Identifying biomarkers could ultimately save lives.
While Platr4 offers one possible route to a better understanding of the human heart, there are others in play as well. One such avenue of exploration is being taken by another member of Spector’s lab.
The science led us there. The findings were a big surprise.”
CSHL Professor David Spector
The second study, published in Nature Communications, also examined gene expression in mESCs and heart cells made from stem cells. However, it looked specifically at alleles in non-sex chromosomes.
Generally, cells have two copies of every gene, called alleles—one from mom and one from dad. In most cases, both copies are active and make RNA. But sometimes, only one is active. This phenomenon is called monoallelic gene expression.

Using RNA sequencing, postdoctoral fellow Gayan Balasooriya was able to identify a significant number of genes that were expressed monoallelically in heart cells. Previously, researchers assumed there were no differences between mom and dad’s alleles in these chromosomes. But Balasooriya also spotted changes that alter gene expression between the paternal and maternal alleles.
“That was an unexpected finding,” Spector says. “That wasn’t known before.”
Next, Balasooriya made another surprising discovery by looking through databases of human and mouse gene expression samples. He found that some males were more susceptible to particular diseases if a critical gene was expressed solely from the single maternal allele.
“The data indicates if a gene is only expressed from the mother’s or father’s allele then that has a potential impact on heart development and health,” Balasooriya says.
While this finding may help researchers understand heart defects, Spector says it may also aid in his lab’s cancer research.
“Not having one allele of a gene turned on can be critical in certain instances,” he explains.
For example, cells in every healthy human body should come equipped with two copies of each tumor-suppressing gene that produces cancer-fighting proteins. But a person with only one copy of the gene turned on would produce half the protein, potentially making them more susceptible to cancer.
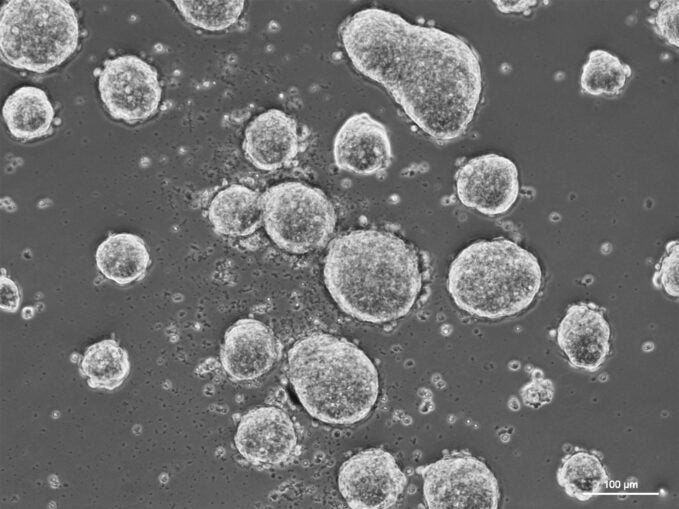
Hazra and Balasooriya are both postdoctoral fellows in the Spector lab. And both have Spector’s support to take their research with them as they continue their careers.
Balasooriya plans to dig deeper into the “fundamental mechanism” of monoallelic gene expression to better understand how it affects the heart. He sees the next step as reassessing the genetic backgrounds of genes already known to cause heart diseases.
“Understanding the fundamentals of maternal and paternal gene regulation may lead to better diagnostics and effective treatment design,” Balasooriya says.
Hazra would like to focus her future studies on the human equivalent of Platr4 and how it affects heart generation. She thinks lncRNAs could target diseases across the board because some of these molecules are only expressed in diseased cells.
Hazra says that “lncRNA-based therapy is gaining a lot of interest. This work can explain why we need to proceed further with this molecule.”
Spector agrees. He says the postdocs’ studies have implications not only for heart disease and cancer but genetics research in general:
“The outcome of both of these papers was that we’ve learned a significant amount about the intricacies of how genes are regulated in cardiac development, either by the expression of one allele or by the use of a long non-coding RNA. And we assume that these processes will occur in other developmental paradigms that could be looked at in the future.”
Written by: Margaret Osborne, Science Writer | publicaffairs@cshl.edu | 516-367-8455
