In the mid-1980s, a rural hamlet called West Valley in upstate New York had a 13-year-old guru named Adam Siepel.
A surprising number of folks knew who he was, or at least as surprising as that can be in a tight-knit community of just over 500 people. Locals would seek him out to teach them, guide them, and solve their problems.
A big part of our motivation is just understanding where we come from.”
The boy did not align chakras or share profound proverbs. Quite simply, he knew how to use a computer. Adults asked him how to set up their computers and install software. His basketball coach even pressed him to create a custom database to record the stats of the varsity team. Siepel was in the seventh grade.
“Adam was the kind of kid you knew would advance with the technology around him,” said Richard DeKay, the now-retired West Valley basketball coach.
Today, Siepel is a Ph.D. and Professor at Cold Spring Harbor Laboratory, where he has directed the Simons Center for Quantitative Biology since 2014.
“What I love about this place is that I’m surrounded by experimentalists who are generating really rich, enormous datasets and asking all sorts of interesting questions that can benefit from the use of computational methods,” Siepel said.
He and his colleagues use sophisticated software programs, data analytics and other computational methods to solve complex biological questions. The Center addresses many of the life sciences, ranging from genomics, neuroscience and plant science, to cancers and other diseases. That collision of big data and biology has created a cross-disciplinary world where computer experts, physicists, statisticians, and biologists work together to unravel biological mysteries.
“In order to really move a field forward, you need to have not only people who have the right skills, but people who are asking the right questions,” he said.
Siepel got his first real taste of quantitative or computational biology when he landed a job at Los Alamos National Laboratory in 1994, right after graduating from Cornell University with a B.S. in Agricultural and Biological Engineering.
At the lab, he and his group applied computational tools and methods to reconstruct the evolutionary histories of HIV. They used the data to chart the epidemiology of the disease and how it spread across the world. The work was a perfect match for Siepel’s computing skills, bioengineering studies, and interest in evolution.
“It was the first time I had encountered this idea of reconstructing the past using mathematical models and computational techniques,” he said. “I just found that fascinating.
Siepel continued to explore and master the different elements that makeup QB after he left Los Alamos in 1996.
At the National Center for Genome Resources, he led the development of ISYS, a software platform that integrated bioinformatics databases, analysis tools, and visualization programs. He worked with algorithms to develop a hypothesis about the evolution of genes and species at the University of New Mexico, earning an M.S. in Computer Science. He focused on computational problems in comparative genomics and created new methods of analysis at the University of California, Santa Cruz, where he received his Ph.D. in Computer Science.
Siepel came full circle in 2006 when he returned to Cornell as an Assistant Professor, eventually building a team that focused on the intersection of genomics, evolutionary biology, statistics, and computer science.
To get a sense of how Siepel and his team at the Simons Center are pushing the field of QB forward, it’s important to understand a quantitative biologist’s role in scientific discovery.
First comes the more classical scientific approaches, like field and experimental work. For the life sciences, this might be crossing different mutant crops, studying a patient’s tumor, recording neural activity in a mouse brain, or discovering archaic human remains.
That last example is pertinent for Siepel, who has refined his computer expertise to answer questions about human evolution.
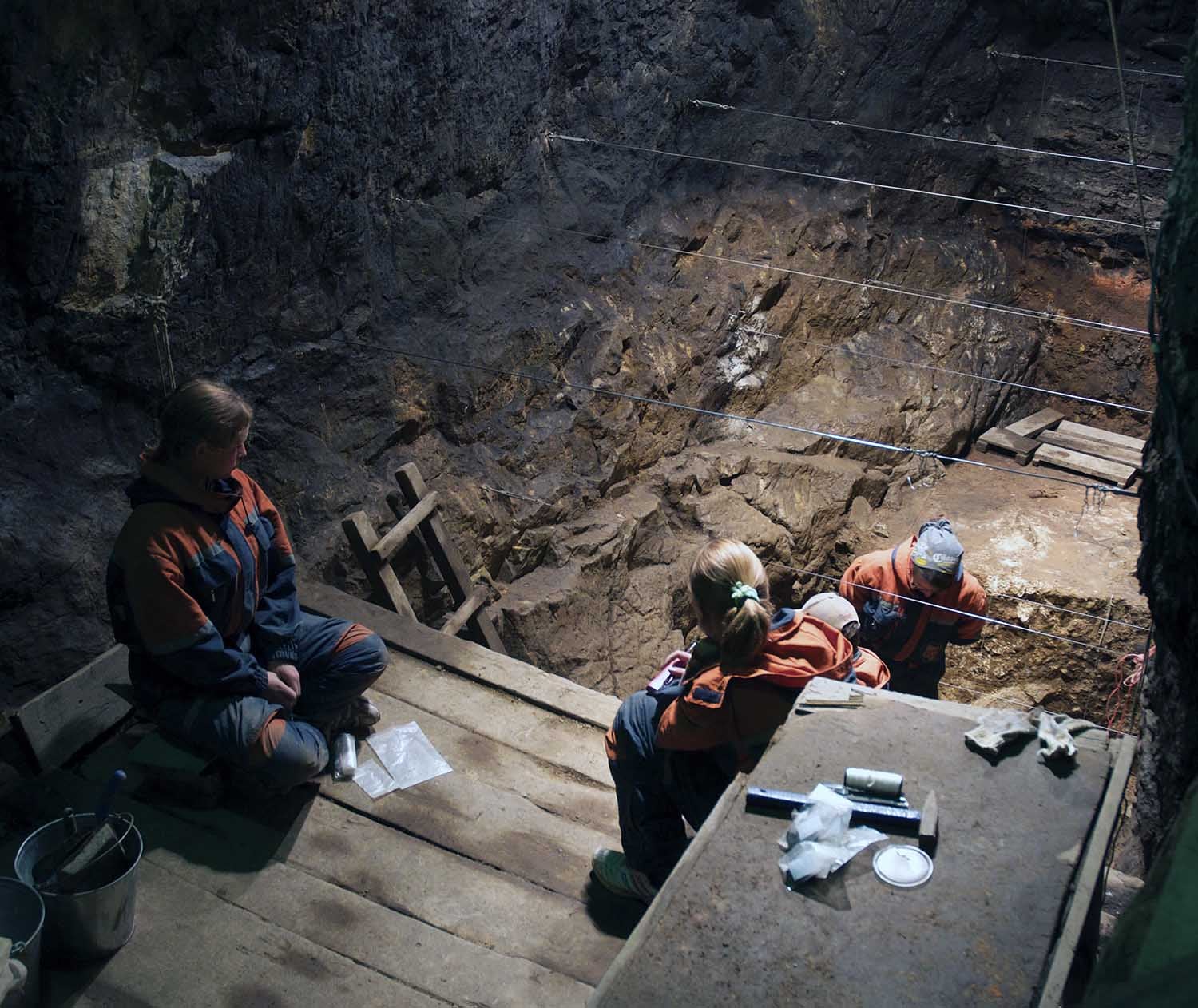

Those questions began shortly after archaeologists and geneticists determined in 2010 that remains discovered in the Denisova cave in Siberia consisted of both Neanderthals and Denisovans, the extinct cousins of modern humans. Scientists gathered whole DNA samples from ancient slivers of bone to determine their age and to which species they belonged. Once those types of conclusions are reached, the question remains: what comes next? Enter the quantitative biologist.
As Siepel sees it, his job is to take many individual answers and bring them together to form an important new observation.
“I’m fascinated by the idea of using powerful mathematical and computation tools to decode the hidden level of meaning in genome sequences,” he said. “The major challenges of biology now are less about measuring things and more about understanding the unifying principles that govern these complex systems.”
In the case of the Neanderthals, Siepel wrote a program to search for certain patterns—such as changes in the individual ancient genomes—that align with the core rules of how evolution alters a genome. In this way, he was able to build a timeline estimating when human populations may have come into contact and interbred with Neanderthal populations. The work, published in 2016, revealed that Neanderthals and humans were swapping genes tens of thousands of years earlier than previously thought.
Why is that important to know? How can a non-scientist benefit from building computer programs that look into our evolutionary past? Siepel gets those questions a lot.
“A big part of our motivation is just understanding where we come from. How did all of what we see in the world come to be? But then, from a more utilitarian perspective, the evolutionary questions are relevant to disease.”
Biologists were able to determine that 1 to 3 percent of the DNA in people with European and East Asian ancestry is of Neanderthal origin, not modern humans. Those Neanderthal portions of the genomes are “enriched for deleterious variants,” meaning that they are more likely to have a negative impact on a person than the rest of their genome. Information like that can lead to the development of disease treatments and prevention.

Combining knowledge like this with computer models that account for the core principles of how cells use DNA allows Siepel and his lab to sift through countless data about which genes are turned on or off, and to search for patterns. Identifying the right patterns puts a cell’s actions, or its expression, into perspective. That could prove essential for diagnosing a disease or finding its cause.
It’s these kinds of discoveries that take computational work way beyond simply crunching numbers.
“I think historically, quantitative biology was basically a field of people who could help analyze data and make sense of large datasets,” Siepel said. “But increasingly, I think it’s possible to actually ask new questions using computational methods.”
Siepel’s own lab includes mathematicians, physicists, and biologists all asking these new questions. It’s an environment where the scientists aren’t thinking in terms of what they study, as you might expect of neuroscientists who study the brain or oncologists who work on cancer. Instead, they use quantitative biology to work on any mystery in the life sciences worth pursuing.
“Every one of our projects involves sort of substantial and deep work from multiple disciplines,” Siepel said. “There is only one science and it’s all connected. With quantitative biology, it’s easy to remember that.”
Written by: Brian Stallard, Content Developer/Communicator | publicaffairs@cshl.edu | 516-367-8455

