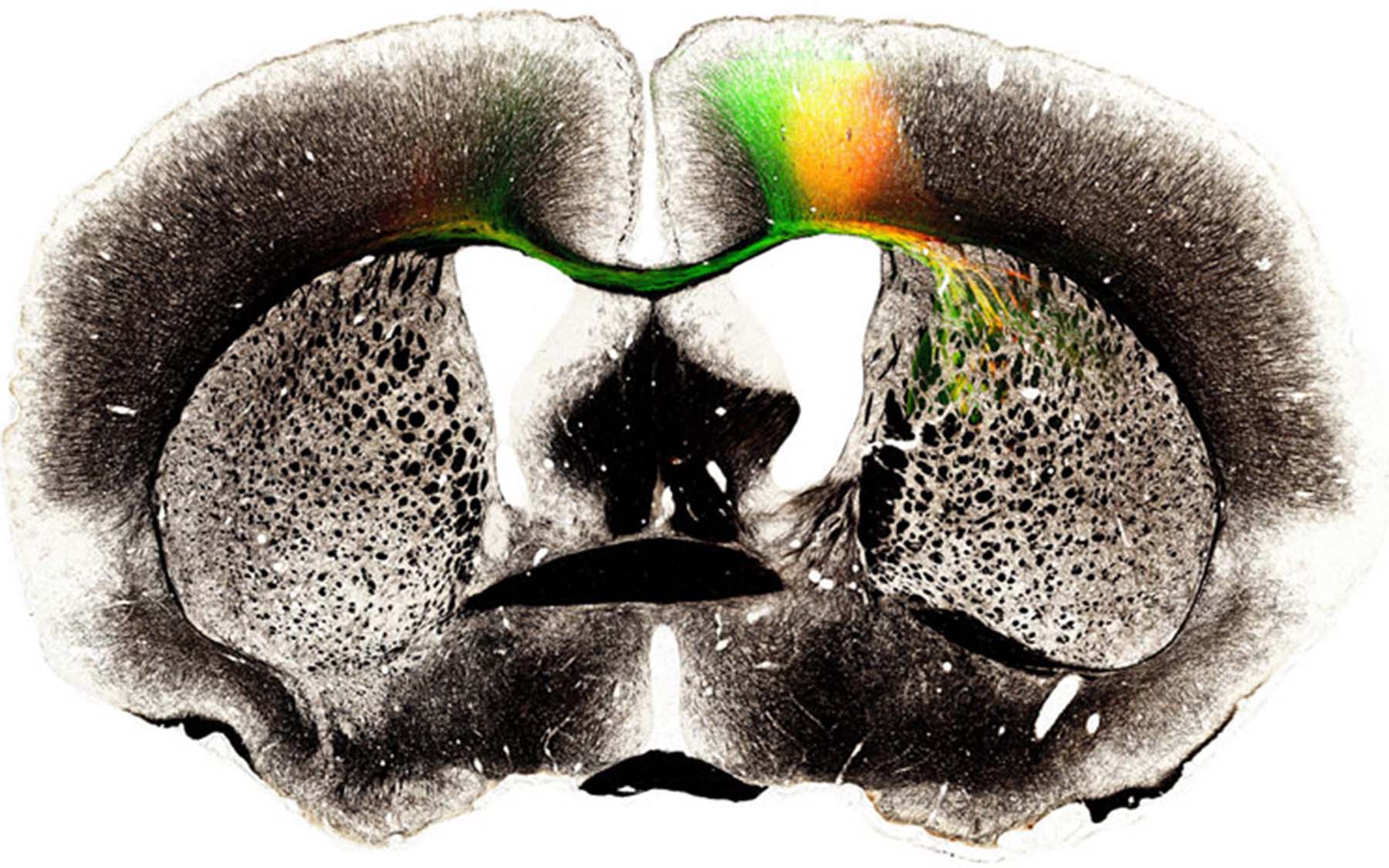
Remarkable imagery enables viewers to journey into the recesses of the mouse brain, following neural connections from end to end.
Cold Spring Harbor, NY — Neuroscientists at Cold Spring Harbor Laboratory (CSHL) reached an important milestone today, publicly releasing the first installment of data from the 500 terabytes so far collected in their pathbreaking project to construct the first whole-brain wiring diagram of a vertebrate brain, that of the mouse.
The data consist of gigapixel images (each close to 1 billion pixels) of whole-brain sections that can be zoomed to show individual neurons and their processes, providing a “virtual microscope.” The images are integrated with other data sources from the web, and are being made fully accessible to neuroscientists as well as interested members of the general public. The data are being released pre-publication in the spirit of open science initiatives that have become familiar in digital astronomy (e.g., Sloan Digital Sky Survey) but are not yet as widespread in neurobiology.
“We’re executing a grid-based “shotgun” strategy for neuronal tract tracing that we first proposed a few years ago, and which I am pleased to note has gained acceptance elsewhere within the neuroscience community,” says Partha P. Mitra, Ph.D., the Crick-Clay Professor of Biomathematics at CSHL and director of the Mouse Brain Architecture (MBA) Project. After the initial June 1 release, project data will be made public continuously on a monthly basis, Mitra says.
Project addresses a large gap in knowledge
“Our project seeks to address a remarkable gap in our knowledge of the brain,” Mitra explains. “Our knowledge of how the brain is wired remains piecemeal and partial after a century of intense activity. Francis Crick and Ted Jones emphasized this in an article published in Nature nearly 20 years ago. Yet to understand how the brain works (or fails to work in neurological or neuropsychiatric disease), it is critical that we understand this wiring diagram more fully. Further, there remain fundamental questions about brain evolution that cannot be addressed without obtaining such wiring diagrams for the brains of different species.”
The MBA Project, which has received critical funding from the Keck Foundation and from the National Institutes of Health, is distinguished by the approach advocated by Mitra and colleagues in a position paper published in 2009. Mitra there proposed mapping vertebrate brains at what he calls the “mesoscopic” scale, a middle-range amenable to light microscopy, providing far more detail than, for instance, MRI-based methods, and yet considerably less detail than is achievable via electron microscopy (EM). The latter approach, while useful for mapping synaptic connections between individual neurons, is feasible on a whole-brain basis only for very small brains (e.g. that of the fruit fly) or very small portions of the mouse brain.
The pragmatic approach Mitra advocated and which is realized in this first data release, is to image whole mouse brains in a semi-automated, quality-controlled process using light microscopy and injected neural tracers (both viruses and classically used tracer substances). While the basic methodology has been available for some time, systematically applying it to a grid of locations spanning the entire brain, and digitizing and re-assembling the resulting collection of brains, is a new approach made feasible by the rapidly falling costs of computer storage. A single mouse brain at light-microscope resolution produces about a terabyte (1 trillion bytes, or 1000 GB) of data; thus, generating and storing the data set currently being gathered would have been prohibitively expensive a decade or so ago.
Assembling the circuit diagram at a mesoscopic scale using ‘shotgun approach’
A key point is that at the mesoscopic scale, the team expects to assemble a picture of connections that are stereotypical—that is, essentially the same in different individuals, and probably genetically determined in a species-specific manner. By dividing the volume of a hemisphere of the mouse brain into 250 equidistant, predefined grid-points, and administering four different kinds of tracer injections at each grid point—in different animals of the same sex and age—the eight-member team at CSHL assisted by collaborating scientists at Boston University, MIT and the University of California, San Diego seeks to assemble a complete wiring diagram that will be stitched together from the full dataset.
The project in this sense is analogous to the Human Genome Project’s “shotgun” approach, in that its final product—a comprehensive wiring diagram—will be the product of many individually obtained data components, woven together thanks to the power of advanced computing and informatics. Indeed, Mitra says one of the genome project’s early advocates, Dr. James D. Watson (now CSHL Chancellor Emeritus), provided him with motivation and encouragement to pursue the project.
“We will never understand how the brain works until we have the wiring diagram,” Dr. Watson comments today. “Mitra is on the right track and I’m impressed he’s gone from conception to putting out data in a couple of years on a quite modest budget. His approach deserves strong funding support.”
The MBA Project was also inspired by early efforts of the Allen Institute, funded by Microsoft co-founder and philanthropist Paul Allen, which resulted in assembly of a comprehensive map of gene expression across the mouse brain. That effort was the product of standard molecular biology procedures iterated in a quasi-industrialized process. The resulting whole-brain gene-expression map, while a triumph, was not designed to shed light on connections in the brain, which became a point of departure for Mitra.
Since the 2009 publication of Mitra and colleagues’ proposal for meso-scale circuit-mapping projects for whole vertebrate brains, the approach has not only spawned Mitra’s CSHL project, but also other meso-scale circuit-mapping projects for the mouse at the Allen Institute and at UCLA. Each differs in aim and technical detail.
A number of features distinguish the “meso-scale” circuit project at CSHL. The 20-micron spacing between brain “slices” gives the CSHL results a particularly rich sense of three-dimensional depth and detail. The team’s use of four tracers including both classical tracer substances as well as neurotropic viruses (attenuated or disabled viruses that infect nerve cells), provides redundancy and helps control for differing efficacies of the different tracer substances. The images one sees on the MBA Project website beginning today provide hard data on actual neuronal processes—the “ground truth” of neuroanatomy, in Mitra’s words—and do not rely on inferential methodologies such as functional MRI scans and diffusion tensor imaging to suggest areas in which connections occur. Finally, it is noteworthy that the slides generated by the project are being physically stored, to permit re-examination at a later date, using more refined imaging methods if necessary or as new methods become available.
“Our project is what I’d call a necessary first step in a much larger enterprise, that of understanding both structure and dynamics of the vertebrate, and ultimately, the human brain,” says Mitra. “While facile comparisons with Genome projects should be avoided, the data sets generated by the MBA and similar projects will provide a useful framework—not unlike a reference genome—on which we can ‘hang’ all kinds of neuroscience knowledge, the body of which has always been notably fragmentary.”
The planning stage of the project, including meetings at the Banbury conference center at CSHL as well as initial informatics work, was made possible by an award from the Keck Foundation.
Written by: Communications Department | publicaffairs@cshl.edu | 516-367-8455
Funding
Major funding for the Mouse Brain Architecture project comes from a Challenge Grant from the National Institutes of Health (RC1MH088659) and a Transformative Award from the Office of the NIH Director (R01MH087988). Additional sources of funding include internal funding at Cold Spring Harbor Laboratory and the Crick-Clay Professorship.
Citation
A technical whitepaper on the project, including a summary of the pipeline used in gathering and processing the data, is available at: http://brainarchitecture.org/mouse/documentation/project-white-paper
2009 position paper proposing the project:
http://www.ploscompbiol.org/article/info:doi/10.1371/journal.pcbi.1000334