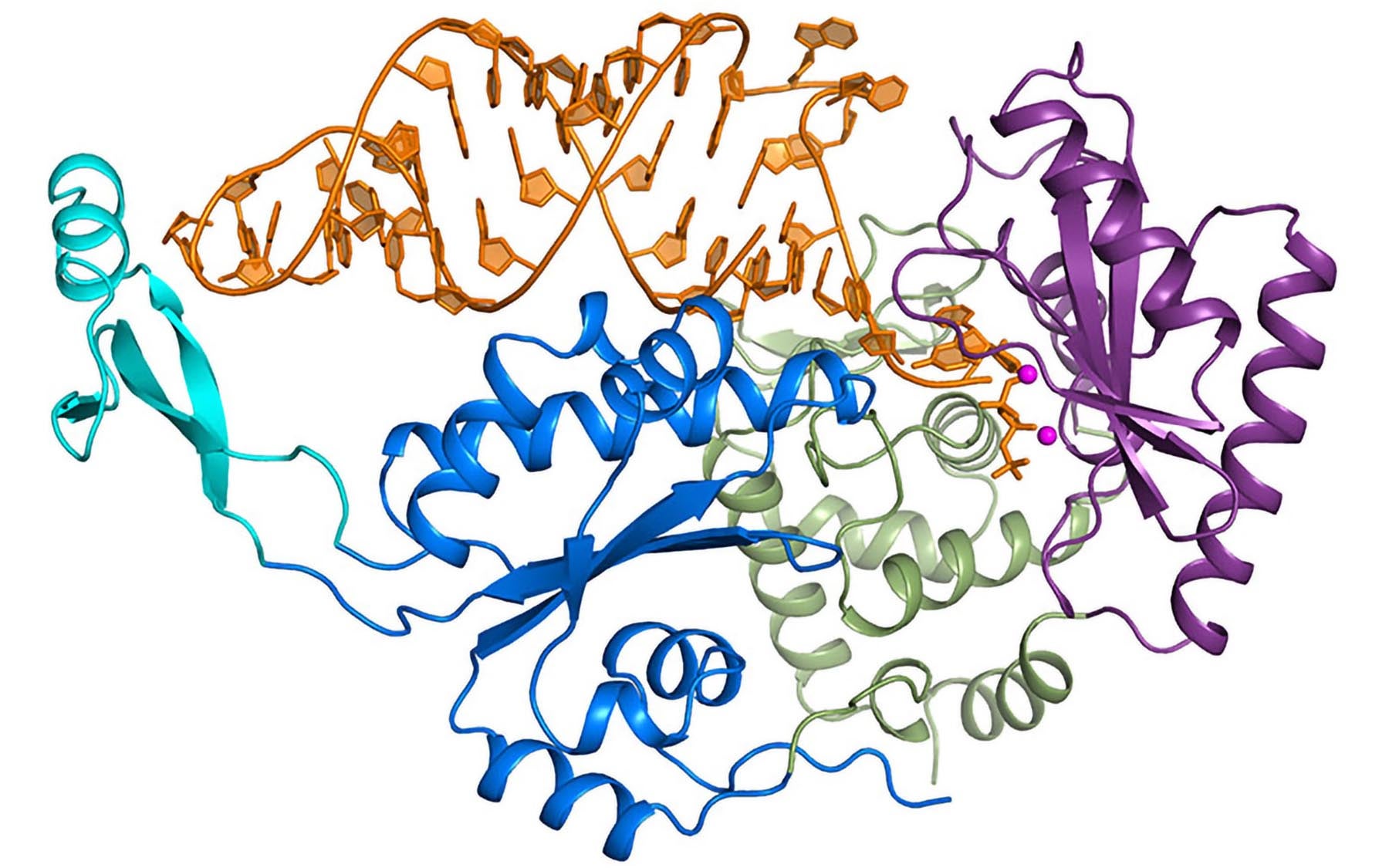
Molecular photographs of an enzyme bound to RNA reveal a new, inherent quality control mechanism
Cold Spring Harbor, NY — Building a protein is a lot like a game of telephone: information is passed along from one messenger to another, creating the potential for errors every step of the way. There are separate, specialized enzymatic machines that proofread at each step, ensuring that the instructions encoded in our DNA are faithfully translated into proteins. Scientists at Cold Spring Harbor Laboratory (CSHL) have uncovered a new quality control mechanism along this path, but in a remarkable role reversal, the proofreading isn’t done by an enzyme. Instead, one of the messengers itself has a built-in mechanism to prevent errors along the way.
The building blocks for proteins are carried by molecules known as transfer RNAs (tRNAs). tRNAs work with other cellular machinery to ensure that the building blocks—amino acids—are arranged in the proper order. But before a building block can be loaded onto a tRNA molecule, a three-part chemical sequence that scientists call “CCA” must be added to the tRNA. The letters are added by an appropriately named machine, the CCA-adding enzyme, and they mark the tRNA as a fully functional molecule.
If a tRNA is mutated, the CCA-adding enzyme duplicates its message. The letters now read “CCACCA,” signaling that the tRNA is flawed. The cell rapidly degrades the aberrant tRNA, preventing the flawed message from propagating.
But how does the CCA-adding enzyme distinguish between normal and mutant tRNAs?
CSHL Professor and Howard Hughes Medical Institute Investigator Leemor Joshua-Tor led a team of researchers to investigate how the CCA-adding enzyme makes this distinction. “We used X-ray crystallography—a type of molecular photography—to observe the enzyme at work, and we were surprised to find that the enzyme doesn’t discriminate at all,” explains Joshua-Tor. “In fact, it is the RNA that is responsible for proofreading itself.”
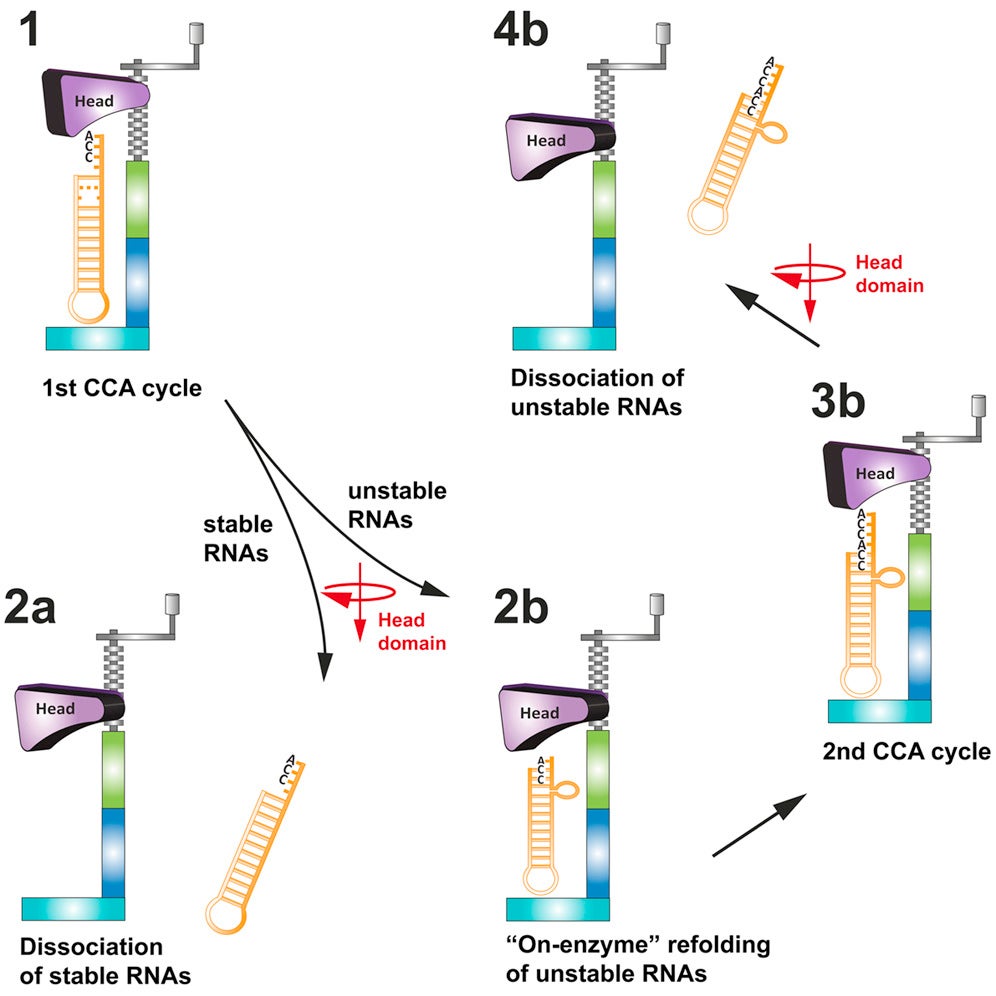
The team used two tRNA-like molecules, called noncoding RNAs, to study the error-correcting mechanism. In previous work, Jeremy Wilusz, PhD, a former CSHL Watson School of Biological Sciences graduate student and an author on this current publication, found a noncoding RNA that is modified with a single CCA group, making it both stable and abundant. Another RNA used in the current study is normally present at negligible levels in cells, and Wilusz and CSHL Professor David Spector found that it is modified with a CCACCA sequence and is rapidly degraded. The difference between the two noncoding RNAs is a simple mutation, and the question the team addressed is how the presence of the mutation affects the addition of “CCA” sequences.
In work published online today in Cell, the team describes a series of molecular photographs of the CCA-adding enzyme bound to the noncoding RNAs. “The CCA-adding enzyme uses a screw-like motion to add each letter of the CCA group to the end of the RNA,” says Claus Kuhn, Ph.D., lead author on the paper. “Under normal circumstances, after the addition of the final letter A, the enzyme tries to ‘turn’ the molecule again, but can’t.” That increased pressure forces the RNA to pop out of its union with the enzyme—with only a single CCA group attached.
But when an RNA is mutated, the researchers found, the structure becomes more flexible. After a single CCA addition, the mutation allows the RNA to buckle under increased pressure. “That bulge allows the enzyme to add an additional round of “CCA” letters, and only then does the RNA pop out,” says Joshua-Tor.
This is a very unique proofreading mechanism, according to Joshua-Tor. “For the enzyme, there is no difference between the two RNAs—it adds CCA in this screw-like motion regardless of what the sequence is. So it is a mutation in the RNA itself that prevent future errors,” ensuring that proteins are made correctly.
Written by: Jaclyn Jansen, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
This work was supported by grants from the US National Institutes of Health, the Howard Hughes Medical Institute, a postdoctoral fellowship from the Jane Coffin Childs Memorial Fund for Medical Research, the Robertson Research Fund of Cold Spring Harbor Laboratory and the Cold Spring Harbor Laboratory Women in Science Award.
Citation
“On-Enzyme Refolding Permits Small RNA and tRNA Surveillance by the CCA-Adding Enzyme” appears online in Cell on January 29, 2015. The authors are: Claus-D. Kuhn, Jeremy Wilusz, Yuxuan Zheng, Peter Beal, and Leemor Joshua-Tor. The paper can be obtained online at: http://www.cell.com/cell/newarticles
Principal Investigator

Leemor Joshua-Tor
Professor, Director of Research & HHMI Investigator
W.M. Keck Professor of Structural Biology
Cancer Center Member
Ph.D., The Weizmann Institute of Science, 1991