A new mathematical model uses gene expression data to predict where neurons are located throughout the brain
Cold Spring Harbor, NY — Throughout the world, there are many different types of people, and their identity can tell a lot about where they live. The type of job they work, the kind of car they drive, and the foods they eat can all be used to predict the country, the state, or maybe even the city a person lives in.
The brain is no different. There are many types of neurons, defined largely by the patterns of genes they use, and they “live” in numerous distinct brain regions. But researchers do not yet have a comprehensive understanding of these neuronal types and how they are distributed in the brain. Today, a team of scientists at Cold Spring Harbor Laboratory (CSHL) led by Professor Partha Mitra describes a new mathematical model that combines large data sets to predict where different types of cells are located within the brain, based on their molecular identity.
Scientists at the Allen Institute for Brain Science in Seattle are using microscopy to directly observe gene activity, one at a time, in razor-thin slices of mouse brain tissue. This approach yields brain maps that are collectively known as the Allen Mouse Brain Atlas. Each individual map shows where a single gene is expressed in the brain. When multiple maps are overlaid, patterns begin to emerge that show how different regions of the brain activate specific and often discrete complements of genes. These patterns are known as “co-expression” profiles.
Elsewhere, other research groups have taken a complementary approach, harvesting a single type of neuron from the brain and profiling all of the genes that are expressed by that cell. But this data lacks the spatial component of the atlas assembled by the Allen Brain Institute.
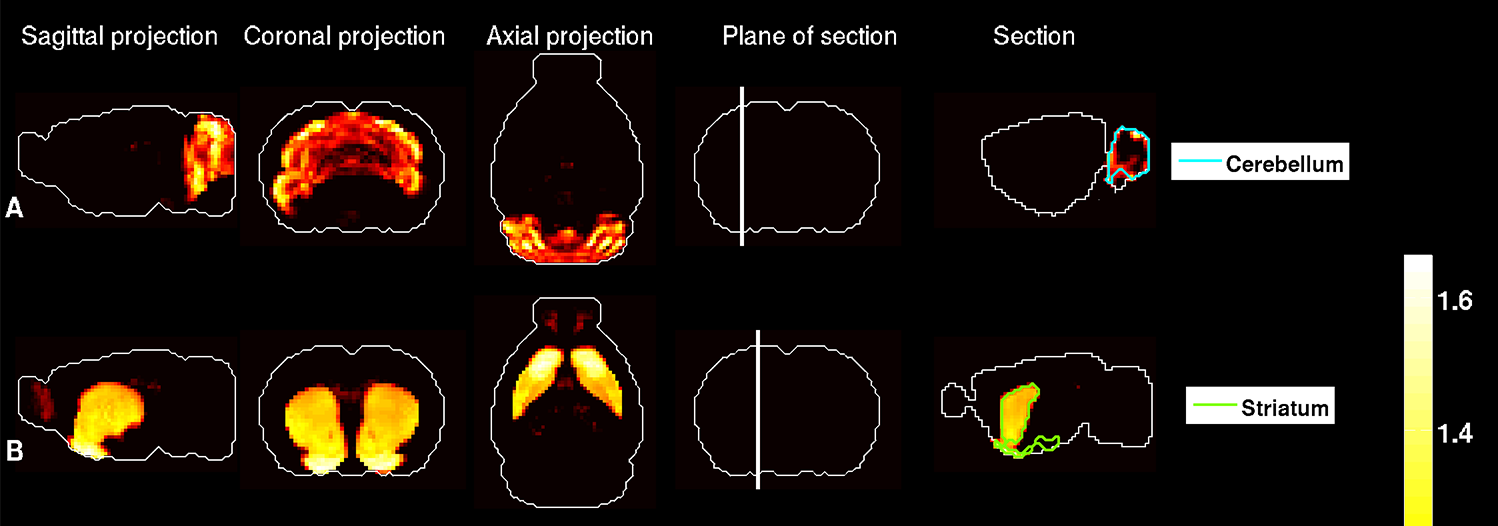
Mitra and postdoctoral fellow Pascal Grange, Ph.D., set out to integrate these two kinds of datasets. They devised a mathematical model that does just this. “Our model is simple,” says Mitra, “but it has predictive power. If the gene expression profile of a neuronal type is measured, then the model predicts where in the brain that type of neuron can be found.”
The significance of the new model, according to Grange, is that “it enables us to now have a biological understanding of the patterns, the co-expression profiles, seen in the Allen Gene Expression Atlas of the Mouse Brain.”
As scientists continue to generate larger datasets of gene activation for neurons, this model will allow them to draw an increasingly accurate map of their distribution in the brain. The eventual goal is to gain a better understanding of how signaling between different types of neurons controls memory and cognition.
Written by: Jaclyn Jansen, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
This work was supported by a grant from the National Institute on Drug Abuse at the National Institute of Health.
Citation
“Cell-type–based model explaining coexpression patterns of genes in the brain” appears online in PNAS on March 24, 2014. The authors are: Pascal Grangea, Jason Bohland, Benjamin Okaty, Ken Sugino, Hemant Bokil, Sacha Nelson, Lydia Ng, Michael Hawrylycz, and Partha Mitra. The paper can be obtained online at: http://www.pnas.org

