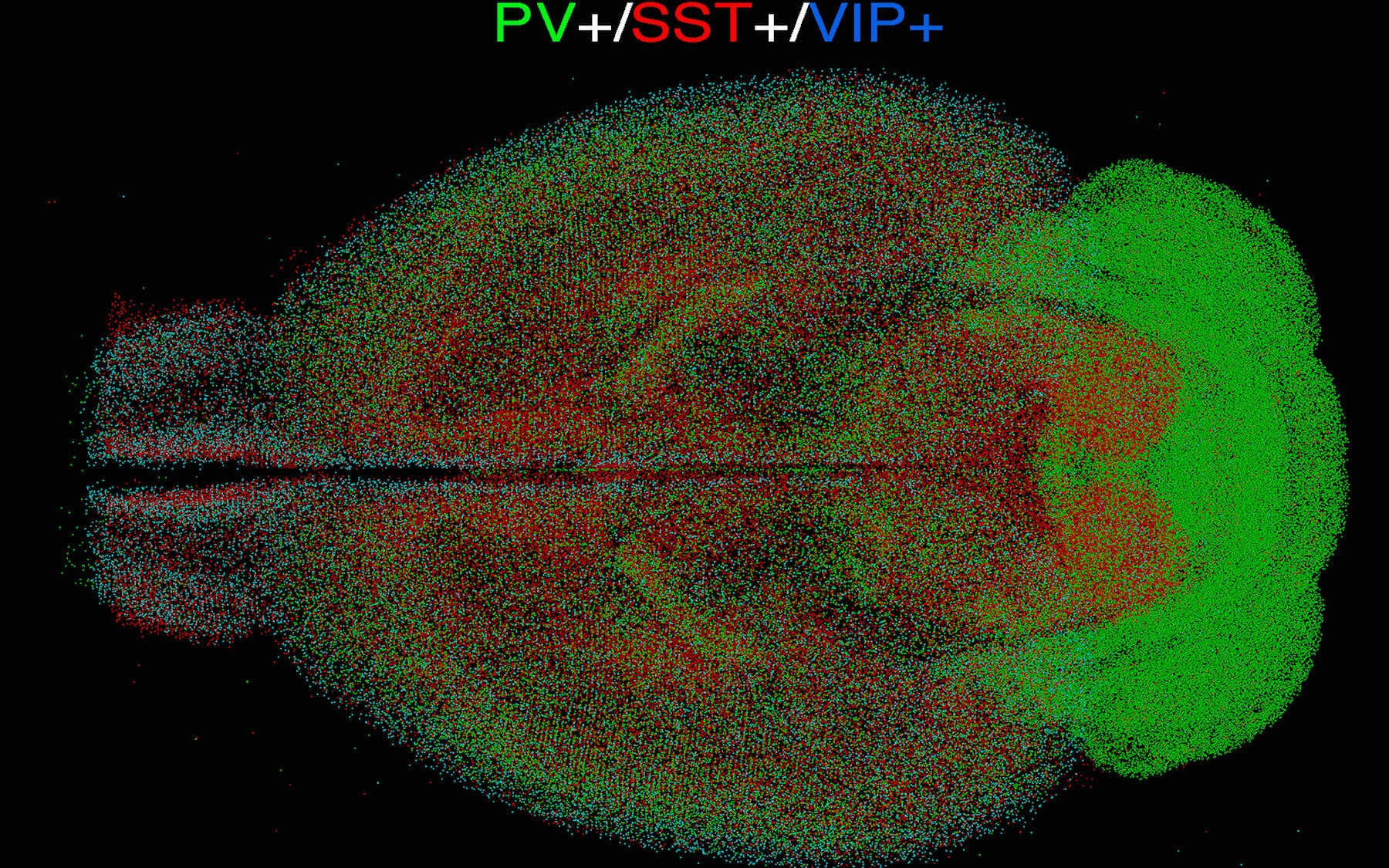
A multiyear project in the Brain Initiative, qBrain is already revealing the brain as never before
Cold Spring Harbor, NY — Neuroscientists at Cold Spring Harbor Laboratory (CSHL) have mobilized advanced imaging and computational methods to comprehensively map—“count”—the total populations of specific types of cells throughout the mouse brain. In a study published today in Cell, they report two highly surprising findings.
Their “qBrain” (quantitative brain) study reveals that contrary to expectations, the numbers and ratios of three major inhibitory cell types vary in a stereotypical way across different parts of the mouse cortex. This implies that different cortical areas, for instance, those involving cognition vs. those involving perception of sensory stimuli, have evolved to tailor their local circuits to specific brain functions. Also, surprising was that while male and female brains did not differ in cell counts in cortical regions, the study identified 11 subcortical areas with sex-specific differences. Strikingly, despite the overall tendency for male brains to be larger, 10 of these regions had more modulatory neurons in females than in males.
“This shows that there are more cells that modulate signals and exert temporal control in areas regulating reproductive, social and parenting behaviors in females than in males, with one exception so far,” says CSHL Associate Professor Pavel Osten, the team leader. The exception is a small area in the hypothalamus, called posterodorsal preoptic nucleus, that is believed to control a single male-specific reproductive function (ejaculation).
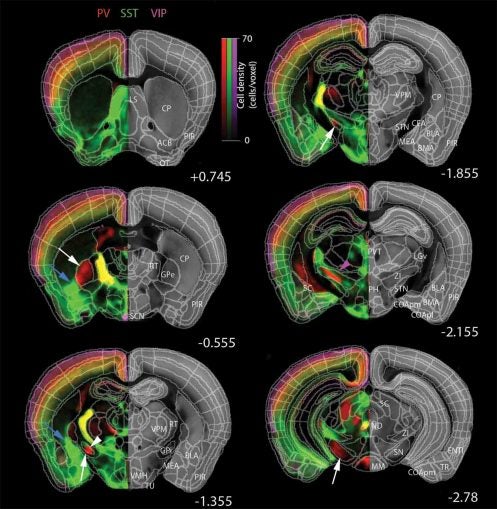
The inhibitory cell types (which modulate the signals of excitatory neurons) counted by the qBrain technology included cells expressing parvalbumin (PV+), somatostatin (SST+) and vasoactive intestinal peptide (VIP+) and four subtypes. Each is a protein that serves to flag a distinct cell type in the mammalian brain. It’s the first step in a project that CSHL and collaborators are performing as part of the National Institute of Health’s “BRAIN Initiative Cell Census Network (BICCN).” In the next 5 years, the team will be building a comprehensive online resource database for the scientific community that will include distribution maps and cellular morphologies for over 100 distinct cell types throughout the mouse brain.
“Even our first results show that it really matters to know the composition of the brain in a precise, quantitative way,” says Osten. qBrain is built on an automated technology platform that will be used to perform similar analyses of other mammalian brains, from prairie voles to marmoset monkeys and humans. This will enable unprecedented cross-species comparisons.
“The brain is like a very complicated Lego puzzle, with pieces that come in all sorts of shapes and sizes,” Osten says. “If you want to understand how brain circuits work, you first need to know how many pieces there are, of what types, and how are they distributed. It’s still not known, but there are thought to be at least 500 cell types in the mammalian brain, and possibly thousands. We still lack a basic parts list.”
CSHL Professors Z. Josh Huang and Partha Mitra collaborated on the qBrain paper. Each mouse brain was scanned and precisely registered in a 600-region grid to a master file by computers, one brain per day. The ~200 GB of data for each brain was then analyzed with machine learning algorithms that identify individual neurons by type, according to parameters “learned” from human experts.
Osten hopes that using qBrain to scan and quantify the brain in mouse models of psychiatric, neurodevelopmental and neurodegenerative disorders—from depression and schizophrenia to autism and Alzheimer’s disease—will provide instructive comparisons with the data drawn from brains of healthy mice. This might enable scientists to begin to clarify what brain irregularities characterize those illnesses.
Written by: Peter Tarr, Senior Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
This research was supported by the NIH, the Simons Foundation Autism Research Initiative, STCSM, the Pennsylvania Department of Health Tobacco CURE Funds, Samuel J. and Joan B. Williamson Fellowship, Mathers Charitable Foundation, HN Mahabala Chair Professorship and IIT Madras.
Citation
Kim Y et al, “Brain-wide Maps Reveal Stereotyped Cell-Type-Based Cortical Architecture and Subcortical Sexual Dimorphism,” appears in Cell October 5, 2017.
Principal Investigator

Pavel Osten
Adjunct Professor
M.D., Medical School of Charles University, Prague, 1991
Ph.D., SUNY Downstate Brooklyn, 1995