“Silent” DNA stays that way as a result of RNA interference during chromosome replication
Cold Spring Harbor, NY — Although letters representing the three billion pairs of molecules that form the “rungs” of the helical DNA “ladder” are routinely called the human “genetic code,” the DNA they comprise transmits traits across generations in a variety of ways, not all of which depend on the sequence of letters in the code.
In some cases, rather than the sequence of “letters,” it is the physical manner in which DNA is spun around protein spools called histones and tightly packed into chromosomes that determines whether or with what intensity specific genes are expressed. A team of scientists at Cold Spring Harbor Laboratory (CSHL) has solved another in a series of mysteries about this critical mechanism of gene expression, described in a paper in the April 8 issue of Current Biology.
Inherited Clumping
According to CSHL professor Rob Martienssen, Ph.D., who led the research team, about a tenth of our DNA stands aloof, spending its time in tightly packed clumps called heterochromatin, and unwinding only to replicate when a cell divides. After copying, both of the resulting DNA molecules—to the surprise of many—have been observed to form reclusive clumps in the same places as the original one did.
This inherited clumping of DNA, which causes genes to be expressed in distinctive ways, is one of a series of phenomena that scientists call epigenetic. The same sequence of nucleotides in two people can produce different patterns of gene expression if the way the DNA is clumped happens to be different.
Probing Epigenetics in Yeast
“We have not understood epigenetic inheritance very well,” says Dr. Martienssen, a plant geneticist and one of the pioneers in the study of epigenetics. To explore this process, he and his team are studying the way DNA is packed in yeast, and how this packing can be transmitted across generations. The single-cell yeast organism is easy to study, in part because it lacks other epigenetic inheritance mechanisms, such as chemical modifications of DNA, that complicate the study of more complex animals and plants.
Long DNA molecules almost miraculously cram into cell nuclei that are almost a million times smaller than they are. They do so by wrapping around proteins called histones, which array themselves along the length of the DNA molecule like beads on a string. These DNA-wrapped histones then form larger arrays. The densely packed mass is then modified chemically by other proteins to form heterochromatin.
The dense packing of heterochromatin hides the DNA sequence from the cellular machinery that reads its genetic information, so the DNA in heterochromatin is “silenced.” The genes it contains are effectively turned off.
Surprisingly, the clumping persists even after cells divide, although, says Dr. Martienssen, “it’s always been a mystery how modifications of histones could be inherited.” A few years ago, however, his group and others solved this mystery. They found that histone modification is controlled by complicated cellular mechanisms broadly known as RNA interference, or RNAi.
In RNAi, RNA that is copied from particular regions of DNA interacts with various proteins to modify histones in the same regions. Because the RNA matches only the section of DNA that produced it, it “provides the specificity that you need to make sure that only that part of the chromosome gets these histone modifications,” Dr. Martienssen says. “If the whole chromosome were to get those histone modifications, you’d be dead.”
All in the Timing
These results raised a new puzzle, though: Since genes contained within heterochromatin are silenced, how can they give rise to the RNA molecules that help to modify histones? In new research, Martienssen’s team has now solved this puzzle by tracking the cells through their cycle of growth and division.
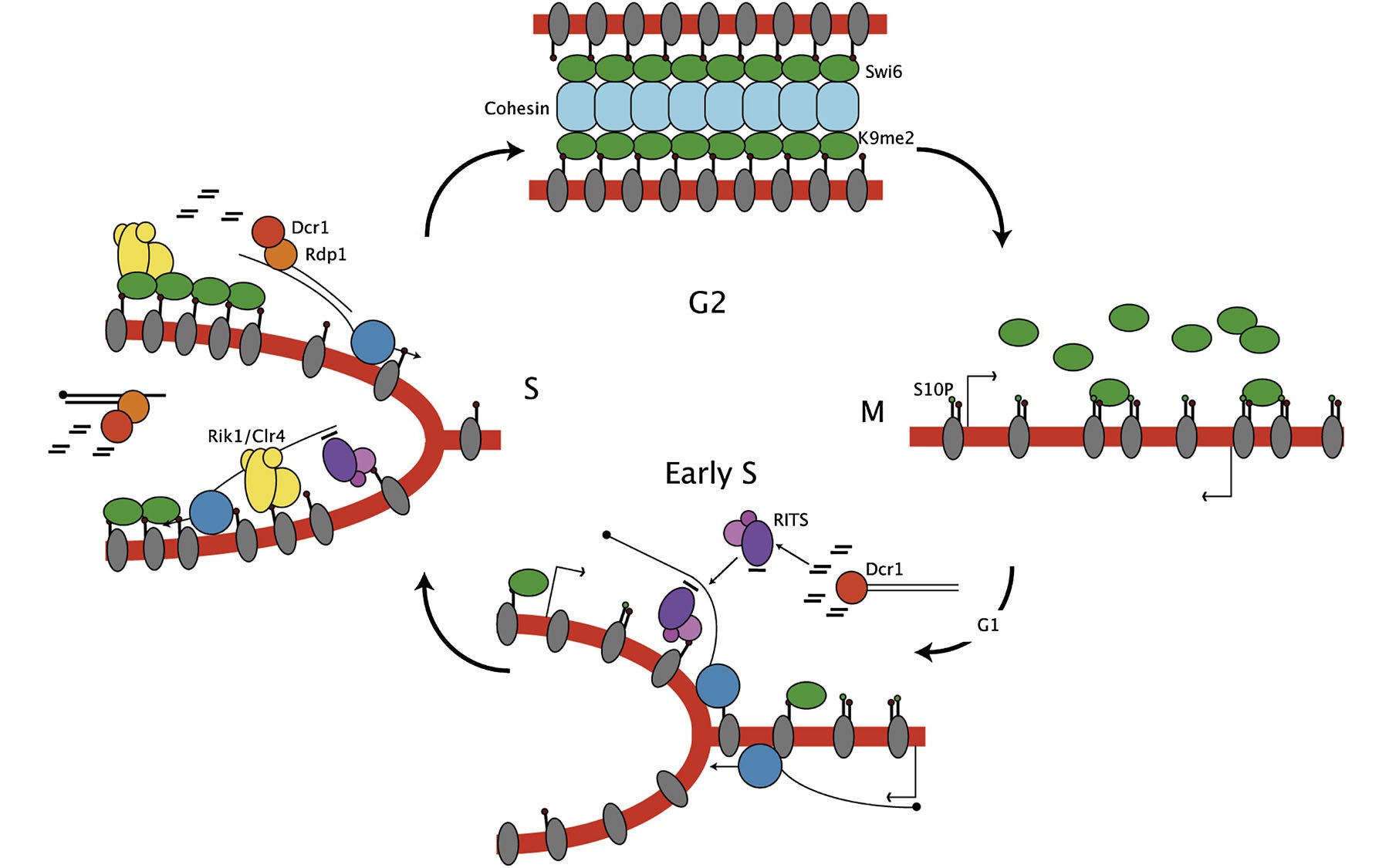
They found that the interfering RNA molecules appear only during the brief part of the cell cycle when DNA is replicating. This result, Martienssen says, “neatly accounts for the paradox about how ‘silent’ heterochromatin can be transcribed [into interfering RNA], because it’s transcribed only in a narrow window of the cell cycle.”
The researchers also found that RNAi varies strongly with temperature. They speculate that this variation is responsible for inherited traits such as vernalization, the well-known process by which certain plants must be exposed to low temperatures before they will flower. Indeed, Martienssen says, there is “a whole slew of epigenetic phenomena that are sensitive to temperature.”
Written by: Communications Department | publicaffairs@cshl.edu | 516-367-8455
Citation
“RNA Interference Guides Histone Modification during the S Phase of Chromosomal Replication” appears in the April 8, 2008, edition of Current Biology. The complete citation is as follows: Anna Kloc, Mikel Zaratiegui, Elphege Nora and Rob Martienssen. The paper is available online at:
http://www.cell.com/current-biology/abstract/S0960-9822(08)00316-3.
Principal Investigator

Rob Martienssen
Professor & HHMI Investigator
William J. Matheson Professor
Cancer Center Member
Ph.D., Cambridge University, 1986