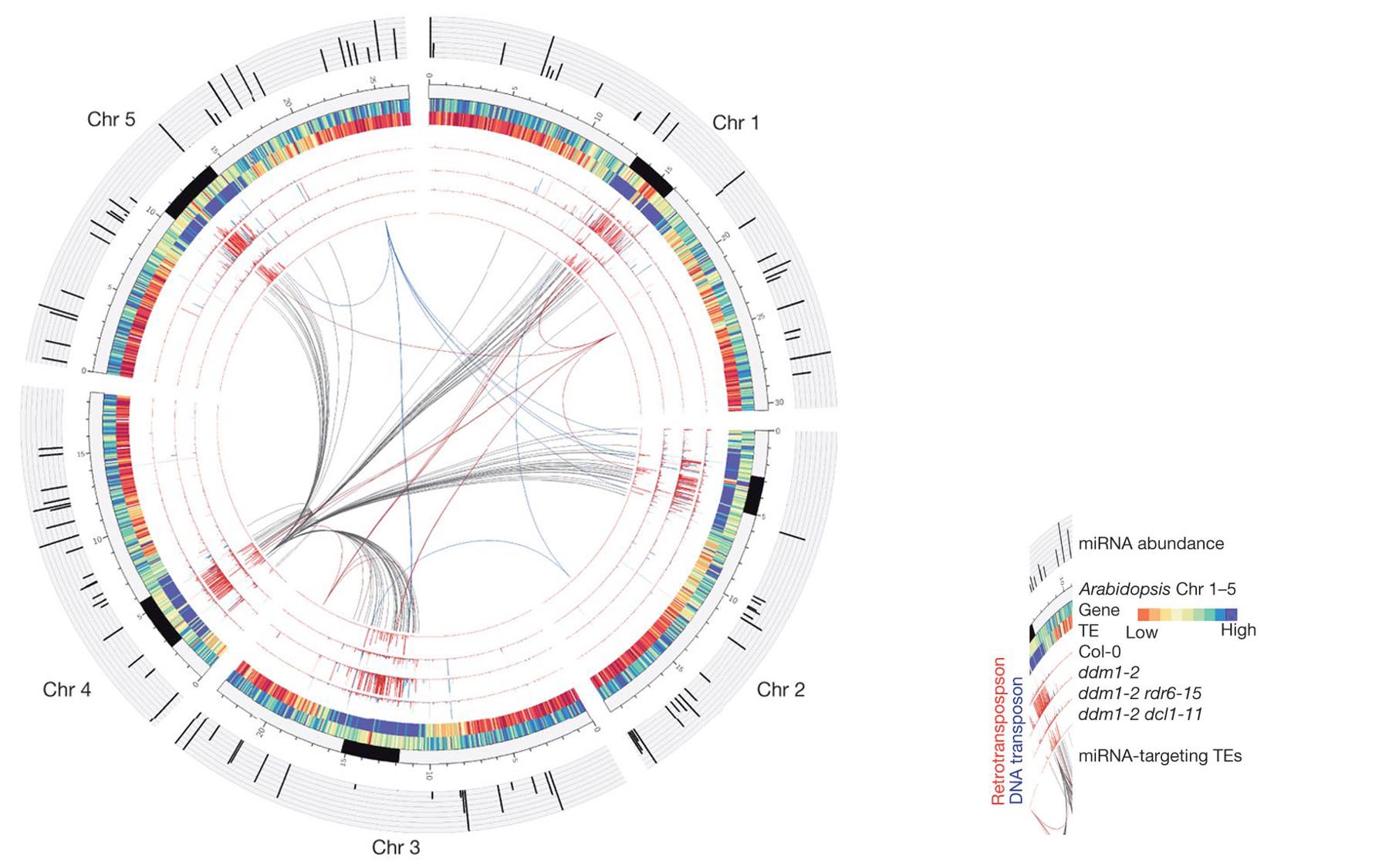
microRNAs are found to target ‘jumping genes’ in reproductive cells, protecting against genomic damage
Cold Spring Harbor, NY — Reproductive cells, such as an egg and sperm, join to form stem cells that can mature into any tissue type. But how do reproductive cells arise? We humans are born with all of the reproductive cells that we will ever produce. But in plants things are very different. They first generate mature, adult cells and only later “reprogram” some of them to produce eggs and sperm.
For a plant to create reproductive cells, it must first erase a key code, a series of tags attached to DNA across the genome known as epigenetic marks. These marks distinguish active and inactive genes. But the marks serve another critical role. They keep a host of damaging transposons, or “jumping genes,” inactive. As the cell wipes away the epigenetic code, it activates transposons, placing the newly formed reproductive cell in great danger of sustaining genetic damage.
Today, researchers at Cold Spring Harbor Laboratory (CSHL) led by Professor and HHMI Investigator Robert Martienssen announce the discovery of a pathway that helps to keep transposons inactive even when the epigenetic code is erased.
“Jumping genes” were first identified more than 50 years ago at CSHL by Nobel Prize-winning researcher Barbara McClintock. Subsequent study revealed that jumping genes (or transposable elements) are long, repetitive stretches of DNA. They resemble remnants of ancient viruses that have inserted themselves into their host DNA. When active, transposons copy themselves and jump around in the genome. They can insert themselves right in the middle of genes, thus interrupting them. Scientists have found that more than 50% of the human genome is made up of transposons. Remarkably, in plants, up to 90% of the genome is composed of these repetitive sequences.

When a transposon is activated, it can insert itself within critical genes, disrupting gene function and causing infertility and many diseases. To combat this ever present threat from within, the cell has devised stringent mechanisms to maintain tight control over transposon activity. The primary mechanism is the epigenetic code, a kind of secondary layer of genetic information that determines how our DNA is used. Epigenetic marks decorate human DNA, delineating active and inactive genes. Regions of the genome that are rich in transposons are heavily marked with inactivating signals, which silence transposons.
The problem for plants, in particular, is that some cells eliminate nearly all epigenetic marks during reproduction. “The loss of these marks puts the cell in tremendous danger, especially at critical times like reproduction,” says CSHL postdoctoral fellow Kate Creasey, Ph.D., lead author on the paper appearing in Nature today. “There must be another mechanism to prevent this kind of widespread genomic disruption.”
The CSHL team discovered a pathway that does precisely this. The pathway they describe acts as a fail-safe to prevent transposon damage when epigenetic silencing is lost. The cell uses small RNAs, known as microRNAs, to accomplish the task. MicroRNAs were already known to regulate gene expression during development. “Now we show that microRNAs actually target transposons when they are activated, for example in the germline (or reproductive cells),” says Martienssen. “This hints that they may have evolved as a transposon defense mechanism.”
Working in collaboration with Professor Blake Meyers at the University of Delaware, Martienssen and his colleagues found that microRNAs silence transposons through a new class of small RNAs known as easiRNAs. Animals have a similar transposon defense mechanism, says Martienssen. “The pathway we have discovered parallels with small RNA systems in animal germlines (called piwi-interacting RNAs, or piRNAs) which also protect against transposons when the genome is being reprogrammed.”
Written by: Jaclyn Jansen, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
This work was supported as part of a collaboration with DuPont Pioneer as well as grants from the National Institutes of Health, the Howard Hughes Medical Institute and Gordon and Betty Moore Foundation, and the Cold Spring Harbor Laboratory Shared Resources by the Cancer Center Support Grant. Additional support was provided by fellowships from the Belgian American Educational Foundation and University of Delaware.
Citation
“miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis” appears online in Nature on March 16, 2014. The authors are: Kate M. Creasey, Jixian Zhai, Filipe Borges, Frederic Van Ex, Michael Regulski, Blake C. Meyers & Robert A. Martienssen. The paper can be obtained online at: http://dx.doi.org/10.1038/nature13069
Principal Investigator

Rob Martienssen
Professor & HHMI Investigator
William J. Matheson Professor
Cancer Center Member
Ph.D., Cambridge University, 1986