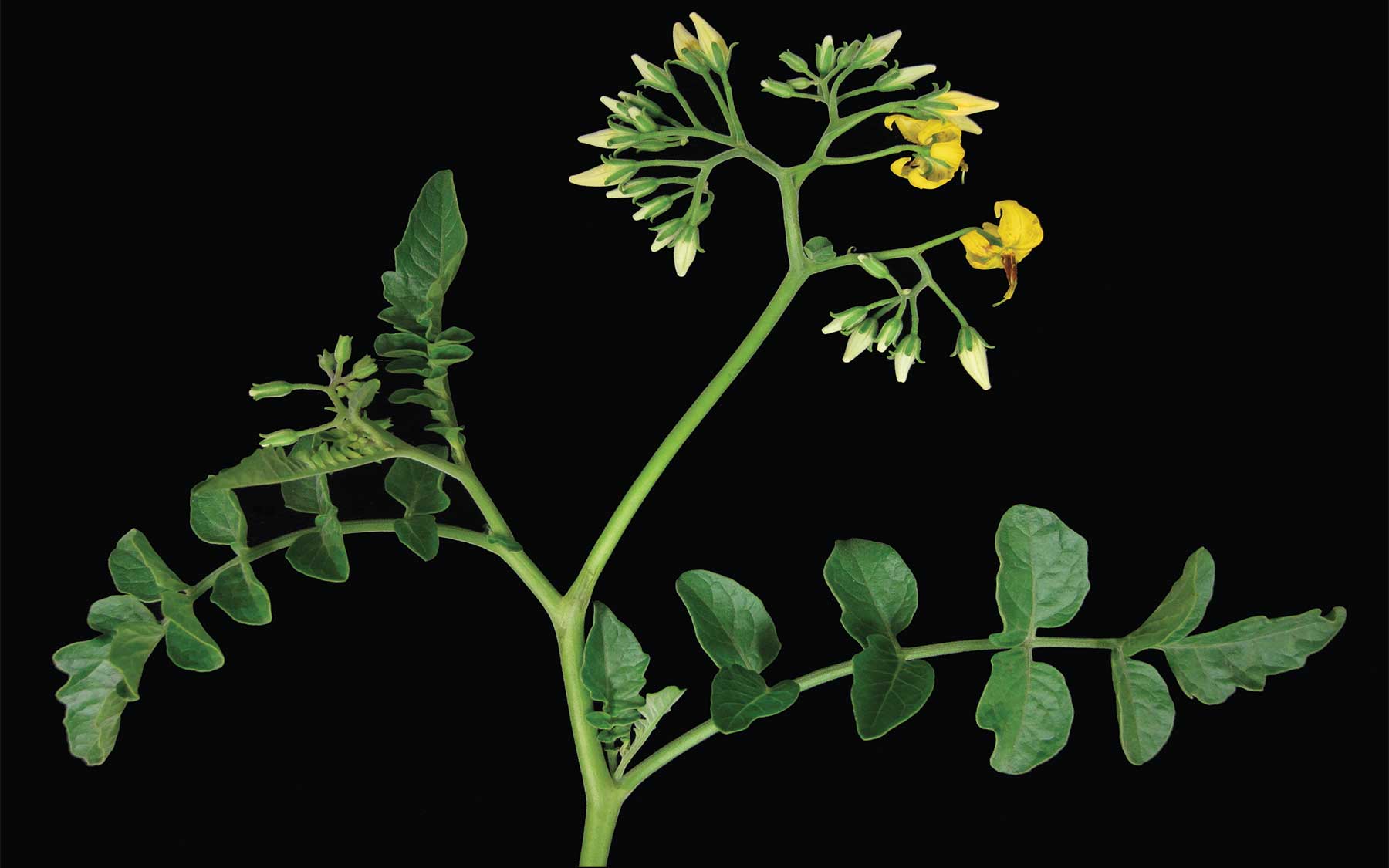
Flower production is directly related to how much our crops yield
Cold Spring Harbor, NY — There is staggering diversity in the number of flowers produced by each of the 2,800 or so species of plants in the nightshade family, which includes economically important crops such as tomatoes, peppers, and potatoes. Some nightshades, such as pepper, produce a single flower, while others, such as tomatoes, give rise to branches bearing multiple flowers.
Why the dramatic differences between such closely related species? While the development of individual flowers is well understood, it has been unclear what mechanisms control how many branches a plant will produce and how many flowers will grow from each branch.
Associate Professor Zachary Lippman and a team of scientists at Cold Spring Harbor Laboratory (CSHL) have used RNA sequencing to identify a network of hundreds of genes that work together to determine the duration of a critical window for the growth of stem cells in plants that give rise to flowers. The longer this window remains open, the more stem cells develop and the more flowers and branches that can grow.

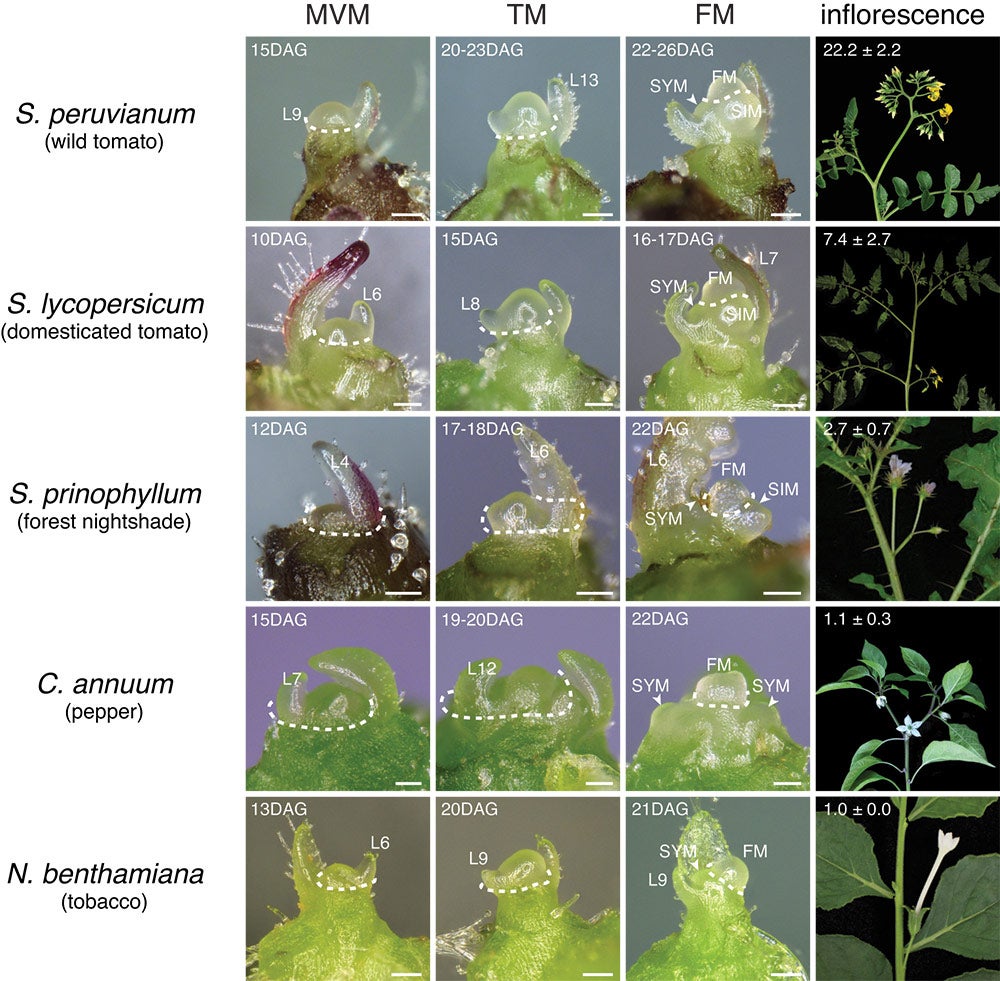
Flower production is an important and widely varying trait in plants. Different plants have different inflorescence architectures, the physical structures that comprise flowers and the branches from which they sprout. Variation can be great even within a single plant family, as shown in these images of 5 members of the Solanaceae family (from l to r): tobacco; pepper; forest nightshade; domesticated tomato; and wild tomato. Their average number of flowers, indicated at the top of each frame, varies dramatically, a quantitative range determined by changes in the stem cell reservoir, or meristem, during the maturation process. (see image below) The longer the maturation, the more complex the inflorescence and the more flowers a given type of plant generates, research from the Lippman lab at CSHL reveals.
The team has also demonstrated that eliminating the activity of three specific genes in this network from tomato plants—genes whose activation usually leads to the production of several flowers—results in plants that produce just one or two flowers.
“Evolution has taken advantage of the flexibility of the plant’s growth schedule in order to create a remarkable diversity in flower production,” says Lippman.
Flower production is enormously important in all crops, serving as the foundation for fruit and seed production. However, significant variation exists between even closely related species, both in the number of flower-producing branching structures, known as inflorescences, and in the number of flowers that sprout from each inflorescence.
Flowers arise from stem cells located at the tips of shoots in dome-like structures called meristems. Until now, the molecular changes in the meristem that determine a plant’s flowering pattern have remained a mystery, primarily because species showing dramatic inflorescence variation, such as the nightshades, are not cross-compatible with one another. This means that traditional crossbreeding experiments cannot be used to pin down the responsible genes.
To overcome this hurdle, Lippman and his colleagues, including CSHL Adjunct Associate Professor Michael Schatz, used RNA sequencing in a comparative genomics approach to look at how the activities of thousands of genes change during the critical window of meristem maturation in five nightshade species.
The team screened nearly 20,000 genes across the five species. Of those, a network of around 300 genes turned on and turned off during the maturation process. The researchers then tracked the activity of those genes during different time periods within the critical window.
As they report today in the journal Genome Research, in plants with simple inflorescences that produce a solitary flower on each inflorescence, a subset of the 300 genes turns on during early stages in the critical window. In plants with more complex inflorescences, the same set of genes turns on a little bit later, extending the window and allowing more time for additional stem cell populations to form. These populations in turn give rise to more branches and more flowers. In a food plant like tomato which produces multi-flowered inflorescences, this results in higher yield.
“There’s a continuum of inflorescence complexity that originates from subtle changes in the maturation schedules in closely related nightshades,” says Lippman.
Though his team studied nightshade plants, Lippman believes the discovery will be a core principle that will explain diversity in different plant families, including the grasses such as corn, rice and wheat. Since these grasses account for many of the world’s leading staple foods, the discovery could have particularly powerful implications.
If evolution has shown it’s possible to find the Goldilocks point in wild plants, we should also be able to achieve the same sweet spot in crops.
— Dr. Zachary Lippman
Researchers wanted to know what specific genes are involved in this process. The nature of the team’s gene expression analysis made it impossible to pinpoint one gene or group of genes. They noticed, however, that among the hundreds of genes in the network was a family of three genes, called BLADE-ON-PETIOLE (BOP), that the team had discovered worked together with a previously studied maturation gene called TMF that ensures tomato plants produce multiple flowers on each inflorescence. These BOP genes have long been known to play a role in plant development, but little had been shown about the importance of these genes in flower production.
As they reported in Genes and Development in September, Lippman’s team used a powerful and precise gene-editing technique called CRISPR to create mutations in each of the BOP genes from a tomato plant. Remarkably, when all three genes were mutated, the inflorescences of the tomato plants produced just one or two flowers.
The BOP genes encode proteins called transcriptional cofactors that control the expression of other genes. Lippman says that these factors likely turn on specific genes during normal growth to delay meristem maturation, essentially extending the critical window in which a plant can produce flowers.
Lippman believes it may be possible to modify the network of 300 genes, including the BOP genes, to manipulate the meristem maturation process, and thereby control the flowering patterns of many types of plants.
“You could tune the maturation process, so to speak, like a dial on a radio for volume control,” says Lippman.
But cranking the dial all the way up to produce as many flowers as possible can backfire, he cautions. A plant that suddenly shifts its resources to creating gobs of flowers will have little energy left to make leaves, fruits and seeds, which are necessary for development and reproduction.
“One nice example of this which we’ve already shown is that we can mutate a gene in tomato plants and get extraordinarily branched inflorescences with hundreds of flowers,” says Lippman. “However, they yield poorly because those flowers just don’t set fruits.”
“If evolution has shown it’s possible to find the Goldilocks point in wild plants,” adds Lippman, “then it means that we should also be able to achieve the same sweet spot in crops, and we believe gene editing is the way forward to achieve this important goal.”
Written by: Chris Palmer, Science Writer | pubaff@cshl.edu | 516-367-8455
Funding
The research described in the release was supported by the National Science Foundation and the Next-Generation BioGreen 21 Program.
Citation
“The Evolution of Inflorescence Diversity in the Nightshades and Heterochrony during Meristem Maturation,” appears online November 7, 2016 in Genome Research. The authors are Zachary H. Lemmon, Soon Ju Park, Ke Jiang, Joyce Van Eck, Michael C. Schatz, and Zachary B. Lippman. The paper can be viewed at http://genome.cshlp.org/content/early/2016/11/07/gr.207837.116.abstract
“Control of inflorescence architecture in tomato by BTB/POZ transcriptional regulators,” appeared online September 15, 2016 in Genes and Development. The authors are Cao Xu, Soon Ju Park, Joyce Van Eck, and Zachary B. Lippman. The paper can be viewed at http://genesdev.cshlp.org/content/30/18/2048.long.
Principal Investigator

Zachary Lippman
Professor & HHMI Investigator
Jacob Goldfield Professor of Genetics
Director of Graduate Studies
Ph.D., Watson School of Biological Sciences at Cold Spring Harbor Laboratory, 2004
