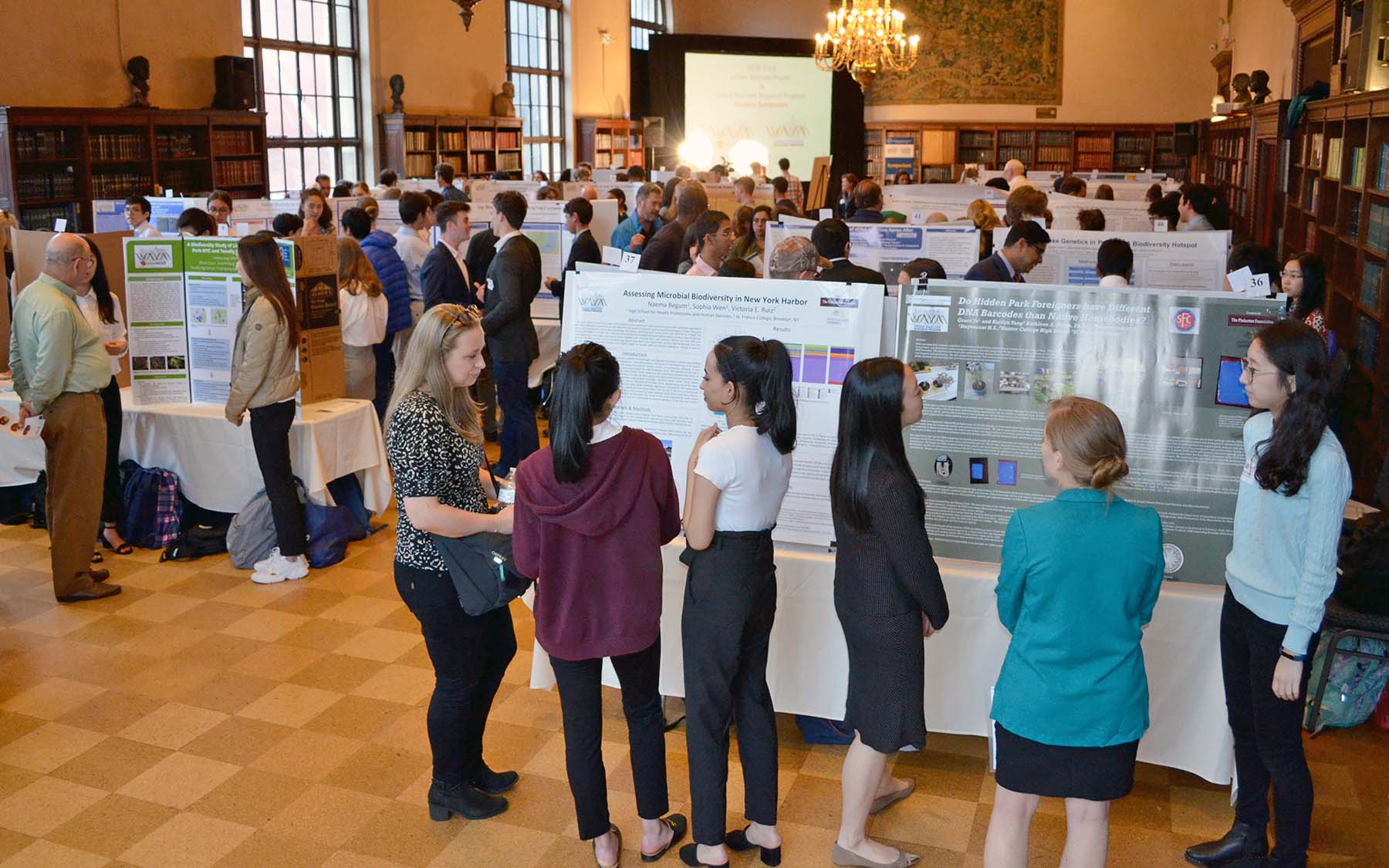
Since 2012, the DNA Learning Center (DNALC) through its unique DNA barcoding programs, has connected science students to research experiences to help them explore urban biodiversity. In the Urban Barcode Project, we demonstrated that large numbers of high school students can work with their classroom teachers to conduct open-ended investigations using DNA barcoding. The Urban Barcode Research Program (UBRP) promotes original research by high school students through collaborations with research institutions across NYC. UBRP students participate in mandatory workshops and conduct independent research projects using DNA barcoding with direct mentorship from scientific experts.
The eighth annual symposium, where students present their projects on biodiversity and engage with science experts, took place May 30th at the New York Academy of Medicine in Manhattan.
More than 160 high school students comprising 61 teams participated in the program and symposium during the last year. The students were mentored by 17 teachers from 13 different high schools, as well as 20 science mentors from 11 academic institutions in the New York metropolitan area.
DNA barcoding started fifteen years ago at CSHL’s Banbury Center where scientists wondered how to use a certain section of DNA to identify an organism in the same way a supermarket scanner uses a barcode to identify a purchase. Turns out that it is indeed possible—the method is effective in identifying each living thing. Over the years, the discipline has evolved, informing major advances in health fields, as well as consumer and environment protection.
By helping to identify species and supplying insight into biological diversity, DNA barcoding provides information that helps shape policies to protect the environment, deal with climate change, improve health regulations, and more. In December 2016, DNA barcoding became a part of the United Nations strategic plan for enhancing and protecting biodiversity.
The student barcoding programs are important science learning opportunities, but the impact of these programs is much broader. While 70 percent of the world’s population is expected to live in urban areas by 2050, urban ecology reviews have criticized the lack of study in urban ecosystems. Cities remain one of the most understudied and poorly understood ecosystems worldwide, demonstrating a huge need for community scientists—like the students in the DNALC programs—to fill the gap.
Students participating in the DNA barcoding programs work side-by-side with trained teachers and expert scientists to learn how specially designed and easily accessible DNA barcoding infrastructure supports a true co-production of knowledge. Several of them took home prestigious awards for outstanding projects or were recognized for their highly relevant studies.
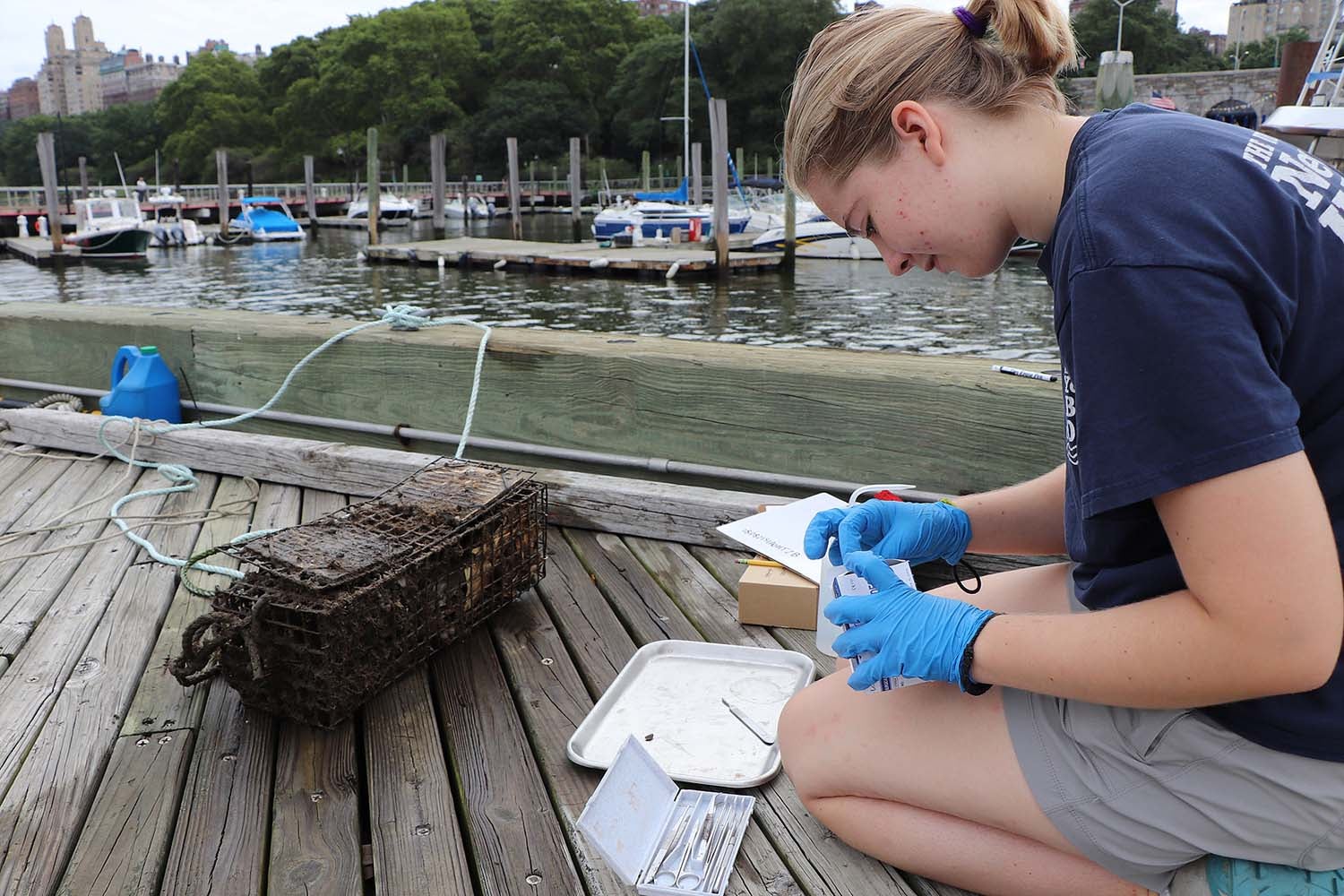

Cyd Bloomfield and Jacqueline Obermayer of New York Harbor School, under the mentorship of Elizabeth Burmester of The Billion Oyster Project, won the International Award for their study on the effects of salinity gradients on biodiversity in the New York Harbor. Bloomfield went on to participate in the New York City Science and Engineering Fair (NYCSEF) and won third place in the earth and environmental sciences category. She also won the American Meteorological Society Award and the Stockholm Junior Water Prize, the world’s most prestigious youth award for a water-related science project.

Damely Abreu and Gissette Noriega of Manhattan Center for Science and Mathematics, mentored by Meagan McMahon of Icahn School of Medicine at Mount Sinai, were recognized for their elaborate virus surveillance study of bird fecal samples in New York City. Wild birds are known to disseminate infectious particles that spread avian diseases and bird-to-bird and bird-to-human transmissions of these diseases are more likely to occur in highly populated areas like NYC. The community scientists understood that assessing the presence of RNA viruses such as Avian Influenza virus and Newcastle Disease virus in NYC birds would help to better prepare for a possible avian disease outbreak, as well as increase the knowledge surrounding avian diseases. The students collected 41 samples from five different city parks, confirmed the birds’ identities by DNA barcoding and tested for the prevalence of both viruses, and even found that a Muscovy duck had detectable NDV, which causes flu-like symptoms in humans.
In another recognized study, Bryce Demopoulos of Hunter College High School and Ethan Lin of Stuyvesant High School worked with mentor Benjamin tenOever of the Icahn School of Medicine to collect 42 invertebrates from their backyards and discovered two novel rhabdoviruses that show high potential to be repurposed for gene editing. The students named the two twin viruses after the Greco-Roman twin deities Castor and Pollux, because they show a high homology.
The students conducted their study because today’s medicine demands the capability to deliver genetic cargo to specific cell types. Past efforts have relied on the engineering of a small subset of vertebrate viruses but have encountered limited success. But as the invertebrate virosphere remains largely unknown, the team sought novel viruses whose components can be characterized and repurposed to build a new suite of viral-based tools.
Thanks to the support of the Pinkerton Foundation, the Thompson Foundation, the Laurie Landeau Foundation, and Science Sandbox, an initiative of the Simons Foundation.
Written by: Christine Marizzi, Manager Student Research Programs at the DNALC | cmarizzi@cshl.edu | 516-367-5170

