The research points to a clear strategy for identifying the subset of liver cancer patients who might benefit from treatment with a therapeutic monoclonal antibody.
Cold Spring Harbor, NY — In a research study appearing in the journal Cancer Cell on March 14, scientists from Cold Spring Harbor Laboratory (CSHL) and four other institutions have identified a strategy for targeted molecular therapy in liver cancer, which currently has limited treatment options and one of the worst one-year survival rates of any cancer type. The researchers’ experiments reveal that up to 15% of liver tumors are “driven” by the hyperactivity of a gene called FGF19, which is well known for its role in various normal biological processes such as cell growth and tissue repair. Shutting down the activity of this gene with an antibody inhibited tumor growth.
“This is an important advance considering that there are currently no genetically-targeted therapies for liver cancer,” says Scott Powers, Ph.D., Director of the Human Cancer Genome Center at CSHL, who led the research team. The new study shows that this gene triggers liver cancer when it exists in multiple copies, well in excess of the normal two per cell. “It turns out that liver tumor cells with this FGF19 amplification are strongly dependent upon the gene’s continued expression to sustain tumor growth.”
This so-called oncogene dependency, in which cancer cells start to rely exceedingly on a single oncogene, is a molecular Achilles heel that cancer researchers are trying to exploit as a strategy for an anti-cancer weapon. This approach has been successful, for instance, by others in targeting breast cancers that have multiple copies of the HER2 gene.
The new study also highlights a powerful strategy that the scientists devised to identify the underlying driver genes that lurk within human cancer “amplicons”—sizeable segments of a cell’s genome that get inadvertently multiplied or amplified, thereby increasing the number of copies of genes that are present within. The strategy, known as oncogenomic cDNA screening, is an extension of an approach developed previously by Powers, Dr. Scott Lowe, and others at CSHL to functionally analyze the human cancer genome.
In the first step of the strategy, the scientists screened tumor samples from 100 liver cancer patients to identify amplicons that occurred frequently among the samples. The 124 genes that exist within the amplicons were then tested for their ability to induce liver cancer when forcibly over-expressed, or hyperactivated, in “sensitized” mice. These are mice that have been genetically engineered to lack a powerful tumor-suppressor protein called p53, while having a hyperactive oncogene called Myc. This deadly combination of genetic errors is seen in more than 40% of human liver cancers.
This experiment revealed 18 of the tested genes to be potent tumor-promoters. The scientists are pursuing studies to understand the mechanisms of how the amplification of these genes result in liver cancer. But from a therapeutic point of view, the most interesting gene was FGF19, as Powers’ collaborator, Dr. Dorothy French at Genentech had previously developed a potent antibody that blocks the activity of the protein produced by this gene.
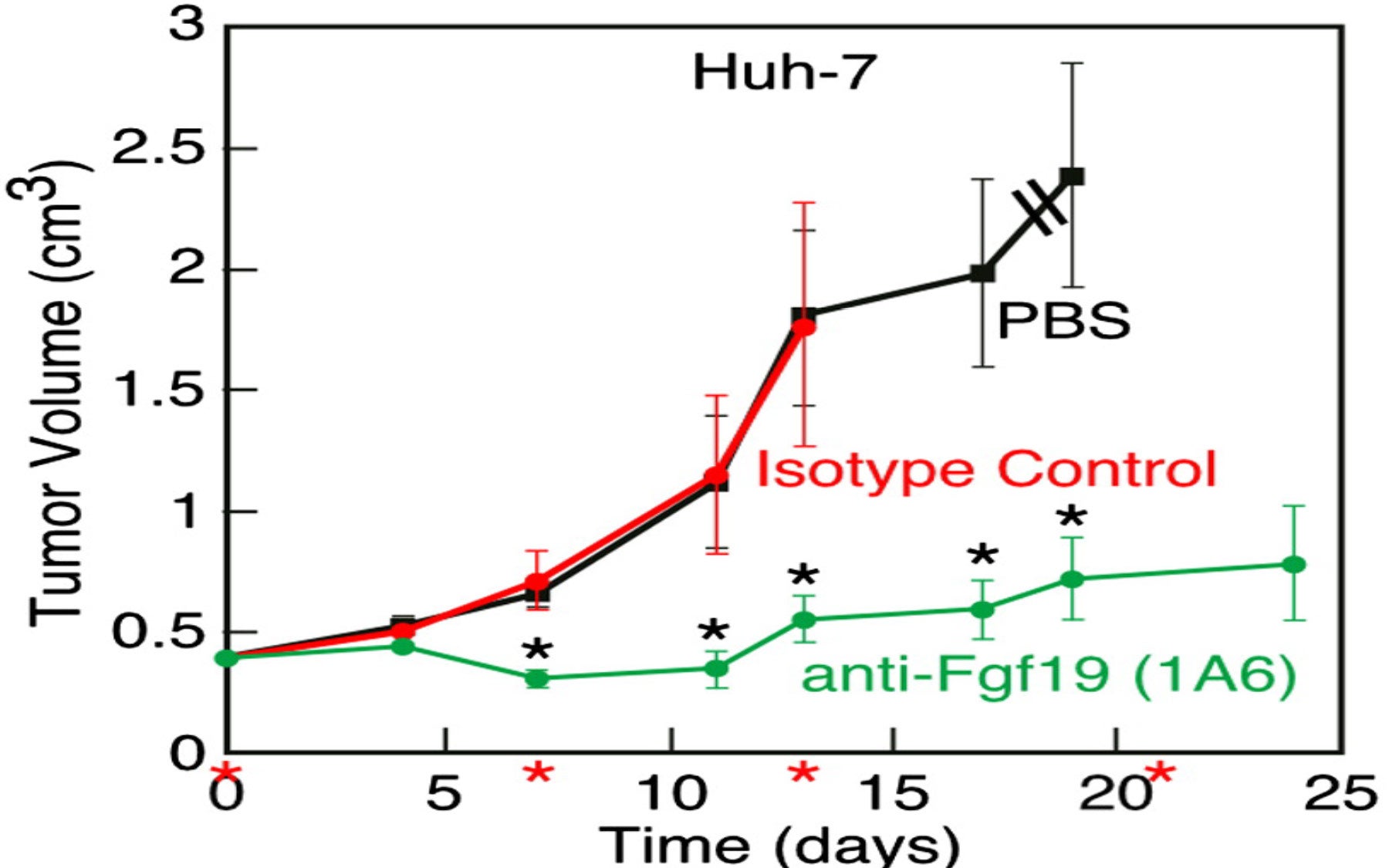
The researchers observed a dramatic inhibitory effect on tumor growth when they injected cancer-bearing mice with this anti-FGF19 antibody. They were also able to significantly suppress tumor growth by shutting down the activity of FGF19 using a technique called RNA interference (RNAi) in which small pieces of RNA that match parts of the RNA produced by the FGF19 gene trigger its destruction.
In other experiments on human liver cancer cell lines, carried out in collaboration with physician-scientist Richard Finn, M.D., at the Jonsson Comprehensive Cancer Center at the University of California in Los Angeles, the anti-FGF19 antibody only suppressed the proliferation of cancer cells that carried multiple copies of FGF19 but had no effect on cancer cells in which this gene had not undergone amplification. “So FGF19 amplification can be used as a biomarker to predict whether treatment with the antibody is going to be effective,” explains first author Eric Sawey, Ph.D., a Research Investigator at CSHL.
Thus, the results not only highlight the potential of using an anti-FGF19 monoclonal antibody as a therapeutic for liver cancer, but also provide a clear strategy for identifying patients who might benefit from this antibody treatment. CSHL scientists are pursuing this effort in collaboration with Dr. Finn who oversees the largest liver cancer clinic in the United States at UCLA.
“Our work represents a new chapter in efforts to expand the current goals of large-scale cancer genome projects that have been unquestionably productive at cataloging genetic changes in cancer,” says Powers. “It shows how oncogenomic screening can leverage this genetic data to pinpoint new therapeutic strategies and drug targets.”
Collaborators on this project also included scientists from the Ontario Institute for Cancer Research and New York University School of Medicine.
Written by: Hema Bashyam, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
The work was supported by the Hope Funds for Cancer Research and grants from the National Institutes of Health.
Citation
“Identification of a Therapeutic Strategy Targeting Amplified FGF19 in Liver Cancer by Oncogenomic Screening” appears online ahead of print on March 14 in Cancer Cell. The full citation is: Eric T. Sawey, Maia Chanrion, Chunlin Cai, Guanming Wu, Jianping Zhang, Lars Zender, Alice Zhao, Ronald W. Busuttil, Herman Yee, Lincoln Stein, Dorothy M. French, Richard S. Finn, Scott W. Lowe, and Scott Powers. The paper can be downloaded at http://www.cell.com/cancer-cell/abstract/S1535-6108(11)00052-3