Sequencing a cancer patient’s “exome” reveals mutations critical for improving diagnostic power and targeted therapy
Cold Spring Harbor, NY — Cancer researchers at Cold Spring Harbor Laboratory (CSHL) have carried out the first comprehensive study of the changes seen in the DNA of a patient with mast cell leukemia (MCL), an extremely aggressive subtype of acute myeloid leukemia (AML) with a very poor prognosis.
Their genomic survey has helped identify two previously unknown mutations that could directly influence patient response to currently available therapeutic drugs.
The details uncovered by the study not only suggest a diagnostic improvement and an alternative treatment strategy for MCL, but could also serve as a springboard for novel diagnostic and therapeutic approaches for other cancers such as lymphoma.
“This is incredibly exciting because we’ve gone from knowing very little about the genetics of MCL to uncovering information that could directly benefit patients diagnosed with MCL,” says Research Investigator Mona Spector, Ph.D., who led the team’s efforts.
The study, which appears online in the journal Leukemia on December 16, is a collaboration between cancer researchers at CSHL and clinicians led by Steven L. Allen, MD, FACP, associate chief of hematology at North Shore-LIJ’s Monter Cancer Center and associate investigator at The Feinstein Institute for Medical Research. “This collaboration between the North Shore-LIJ Health System and CSHL allows us to increase medical knowledge and make innovative discoveries that may lead to new treatments for patients who are living with MCL,” says Allen.
Made possible in large part by funding from the Don Monti Memorial Research Foundation, “the goal of this collaboration was to sequence patient DNA to find information about individual cancers that could be used to improve or design patient-specific treatment strategies,” explains CSHL Adjunct Professor and HHMI Investigator Scott Lowe, Ph.D.
In this study, the CSHL scientists used two approaches to identify genetic changes seen in an MCL patient who succumbed to the disease about three months after diagnosis. MCL is characterized by out-of-control proliferation of transformed mast cells—the same immune system cells that are notorious for their release of histamine during an allergic response.
In one approach, the CSHL team used a method called array comparative genomic hybridization (aCGH) to identify copy number variations—genomic alterations that result in an abnormal number of copies of one or more sections of DNA—in the leukemic mast cells. In a second approach, the team sequenced the majority of the “exome,”—the “exons” which are the DNA sequences that encode for protein (only about 2% of the genome).
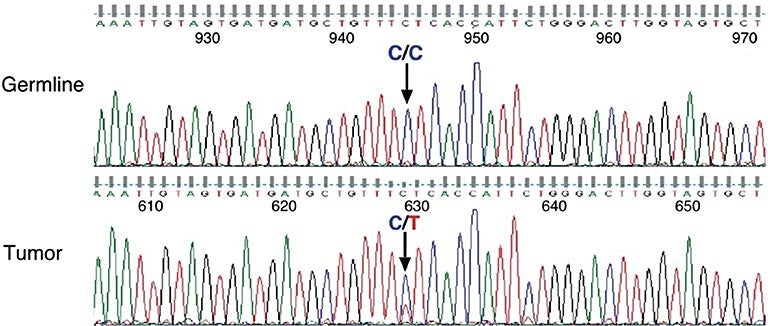
This enabled them to identify mutations that result in the difference of a single nucleotide, or chemical “rung” in the DNA “ladder,” between the patient’s normal and tumor cells. These mutations often result in the production of aberrant proteins and can cause a cell to grow uncontrollably.
Bioinformatic analysis of the gigabytes of sequencing data by CSHL Fellow Ivan Iossifov, Ph.D., revealed the differences between the two genomes. Although several of the mutations occur within genes that have been previously linked to cancer, Spector immediately zeroed in on the mutations in two genes, KIT and MS4A2.
“MCL patients are screened for a mutation in the KIT gene that occurs at a specific amino acid referred to as D816V. This mutation not only spurs uncontrolled mast cell proliferation but also causes resistance to imatinib, a drug that works against some forms of leukemia; so patients with this mutation would normally not be treated with this drug,” explains Spector.
“Our analysis showed that this patient, who lacked D816V and therefore received the drug, actually had a different KIT mutation called V654A, which may also cause resistance to the drug. Had this information been known before, the patient might have been treated differently and been spared the drug’s side-effects.” Spector hopes that this information might now encourage physicians to screen MCL patients for both KIT mutations.
The second mutation of interest occurs within the MS4A2 gene, which encodes for a protein that is part of a receptor that sits on a mast cell’s surface and is required for its survival. The mutation identified by Spector occurs within the region of the protein that is essential both for its presence on the cell’s surface, and more importantly, for triggering intracellular signaling by the enzyme Syk kinase. So Spector suspects that the MS4A2 mutation might be an “activating” mutation that may constantly keep the Syk signal in an “on” state, thereby prolonging the mast cell’s life and eventually leading to cancer.
“If we prove this to be the case, then our finding could be therapeutically exploited because there already is a drug that blocks Syk signaling that has shown efficacy in a clinical trial for lymphoma,” says Spector. She also observes that their findings might have implications beyond MCL. “For example, Syk signaling is also important in another type of immune cell called B cells, so researchers studying B cell cancers might also want to now look for mutations in genes within this pathway to identify patients who might respond to the Syk inhibitor drug,” she says.
Written by: Hema Bashyam, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
The study was supported by funding from the Don Monti Memorial Research Foundation and The Ryan Gibson Foundation.
Citation
“Mast-cell leukemia exome sequencing reveals a mutation in the IgE mast-cell receptor b chain and KIT V654A,” appears online ahead of print in Leukemia on December 16. The full citation is: MS Spector, I Iossifov, A Kritharis, C He, JE Kolitz, SW Lowe and SL Allen. The paper can be downloaded at: http://www.nature.com/leu/journal/vaop/ncurrent/full/leu2011354a.html