The list of short RNAs grows longer
Cold Spring Harbor, NY — RNA is best known as a working copy of the DNA sequence of genes. In this role, it’s a carrier of the genes’ instructions to the cell, which manufactures proteins according to information in the RNA molecule.
But molecular biologists have increasingly realized that many RNA snippets—so-called small RNAs—also directly influence which genes make proteins, and in some cases, how much protein. They’ve also found that some small RNAs play a unique role in protecting the integrity of genetic material.
Scientists at Cold Spring Harbor Laboratory (CSHL) have played a central role in these discoveries since the beginning of the decade. In recent weeks they have published additional findings adding to our knowledge of small RNAs, identifying a brand new class and clarifying how a known class acts to regulate gene activity.
“It turns out that there are more types of small RNA molecules than anyone initially suspected,” said Gregory J. Hannon, Ph.D., CSHL professor and pioneer in small RNA research. “And we are finding that each type that we discover acts in more ways than had previously been appreciated.”
Advanced Sequencing
RNA molecules consist of sequences of chemical units, or “bases,” that are copies of the DNA sequence. Each specific sequence can “recognize” related sequences in other RNA or DNA molecules. To regulate cellular activities, small RNA snippets, each containing 20 to 30 bases, join with special protein helpers to eliminate or modify target molecules. But of the billions of bases in a cell’s DNA, what determines which RNA snippets are chosen for this role?
Dr. Hannon and his collaborators are harnessing highly efficient new machines that determine the sequence of bases in millions of small RNA molecules simultaneously. They then scan the known genome to find matching sequences, as well as the sequences nearby. This original context is crucial to understanding why some snippets are chosen as regulators.
A New Class of RNA
Previously, researchers had recognized two classes of regulatory small RNA molecules in fruit flies, each with different protein partners. One class, called microRNAs, appears throughout the organism and acts to regulate the activity of many genes by joining with a special protein called Argonaute 1. A second class of RNA molecules, called piRNAs, occurs only in cells of the sex organs and joins with different proteins, called Piwi proteins. Together, they act as a kind of immune system to suppress genetic interlopers called transposable elements, which were discovered at CSHL by Nobel laureate Barbara McClintock more than a half-century ago, and which in certain cases can cause genomic havoc that underlies disease.
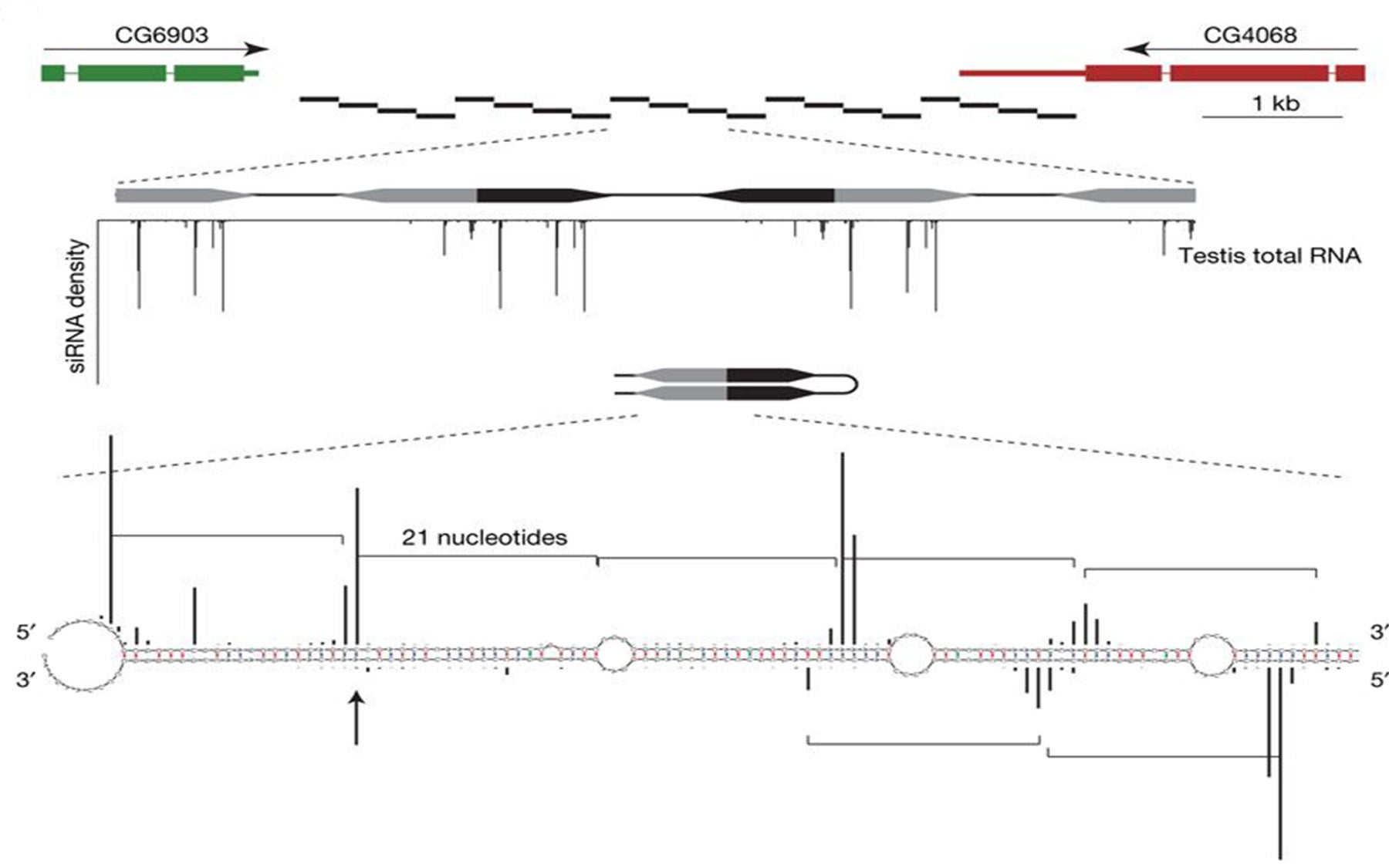
Dr. Hannon and his colleagues looked for RNA molecules that partner with a different protein, Argonaute 2, that belongs to the same family as Argonaute 1 and the Piwi proteins. Using advanced sequencing technology, they found that these small-RNA partners were distinct from either of the previously known classes of small RNAs. The new small-RNA class both modifies gene activity and suppresses transposable elements, thus serving as a defense mechanism.
These findings, the scientists write in a newly published paper in Nature, “expands the known repertoire” of small RNAs in fruit flies, and further blurs distinctions between the previously identified classes of small RNAs.
Pseudogenes: Not Just Junk
In related research, Hannon and his colleagues have published a paper announcing their discovery of a new source of regulatory RNA in mice. Many RNA sequences, such as microRNAs, are flagged as regulatory molecules because they physically fold on themselves. Special proteins recognize the resulting double-stranded RNA, and chemically slice it to release regulatory RNA snippets.
The CSHL team found that double-stranded structures also form from “pseudogenes.” Pseudogenes, in the past assumed to be useless “junk DNA,” are damaged copies of normal genes left over from previous genetic events. The researchers found that RNA copies of normal genes sometimes pair up with copies from the related pseudogenes, resulting in double-stranded RNAs that—far from being junk—are able to activate the cell’s regulatory apparatus.
The new findings add another layer of complexity to our understanding of the byzantine processes by which small RNA molecules influence genetic activity. “Viewed in combination,” Dr. Hannon and colleagues write, “our studies suggest an evolutionarily widespread adoption of double-stranded RNAs as regulatory molecules.”
Written by: Communications Department | publicaffairs@cshl.edu | 516-367-8455
Citation
“An endogenous small interfering RNA pathway in Drosophila” appeared in the advance online edition of Nature on May 7. The complete citation is: Benjamin Czech, Colin D. Malone1, Rui Zhou, Alexander Stark, Catherine Schlingeheyde, Monica Dus, Norbert Perrimon, Manolis Kellis, James A. Wohlschlegel, Ravi Sachidanandam, Gregory J. Hannon, and Julius Brennecke. The paper is available online at http://dx.doi.org/10.1038/nature07007.
“Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes” appears in the April 10, 2008 online edition of the Nature. The complete citation is: Oliver H. Tam, Alexei A. Aravin, Paula Stein, Angelique Girard, Elizabeth P. Murchison, Sihem Cheloufi, Emily Hodges, Martin Anger, Ravi Sachidanandam, Richard M. Schultz, and Gregory J. Hannon. The paper is available online at http://dx.doi.org/10.1038/nature06904.