A Genetic Cancer Genome Project is envisioned to identify new targets for cancer treatments
Cold Spring Harbor, NY — Medicine’s cancer-fighting arsenal is not as accurate as many patients might wish. Chemotherapy and radiation treatments often cause serious side effects because they damage normal cells as well as cancerous ones.
Scientists at Cold Spring Harbor Laboratory (CSHL) are part of a team that has announced the development of a faster and more affordable method for discovering genes essential to the survival of cancer cells—genes that can be suppressed without harming normal cells. Other team members are from Harvard Medical School and Massachusetts General Hospital.
Scanning For Cancer Proliferation Genes Across the Genome
An enhanced capability of isolating such genes, called cancer proliferation and survival genes, is likely to spur new searches for them across the human genome, a process that has the potential to revolutionize medicine’s approach to cancer treatment.
The new method of isolating proliferation genes involves injecting into cancerous cells small strands of RNA called short hairpin RNAs. These can target a specific gene and stop it from being expressed. If the cancerous cells die off, and the same short-hairpin RNA doesn’t kill off healthy cells, then the gene it targets is a cancer proliferation and survival gene.
Previously, researchers typically investigated one gene at a time and couldn’t accommodate a genome-wide search. In order to investigate many genes at once, the CSHL team and their colleagues in Boston have developed an inventory of more efficient short hairpin RNAs—called micro-RNA-based short hairpin RNA libraries—and better markers to track their survival.
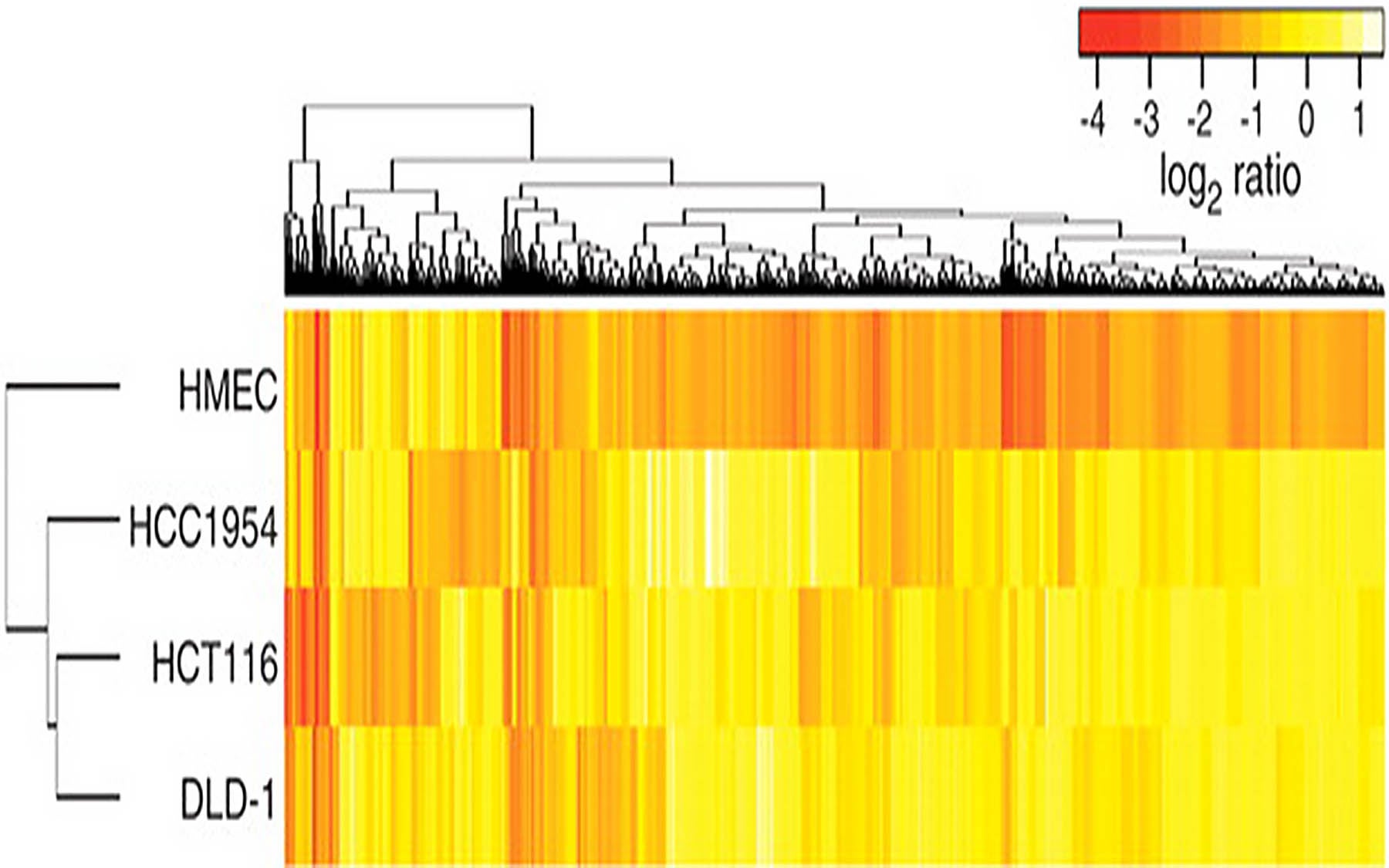
“Until now, the genome-wide search for cancer proliferation and survival genes could only be performed by so-called core facilities in major institutions that had spent several millions dollars in equipment and reagents,” said Gregory Hannon, Ph.D. CSHL professor, Howard Hughes Medical Institute investigator, and co-author of a paper published by the team in the February 1 issue of Science. An accompanying paper published in the same issue, in which Dr. Hannon collaborated with Jose Silva, Ph.D. and others at CSHL and at Harvard Medical School, reports how essential genes in human mammary tissue were profiled using the technique, called multiplex RNAi screening, developed in the Hannon lab.
“Our approach makes it possible for individual laboratories to do these studies for less than 1 percent of the cost and without a dependency on expensive robotics,” Dr. Hannon observed. “The difference in cost and the absence of special equipment gives the unique opportunity to more laboratories of different financial means or size to do these types of studies in only a matter of weeks.”
Toward a ‘Genetic Cancer Genome Project’
The advantages of the new method have prompted Dr. Hannon and his co-authors to propose a “Genetic Cancer Genome Project” in which geneticists will identify and catalog cancer type-specific genes with the goal of revealing potential drug targets.
This could mean that the next generation of cancer treatments will include fewer side effects, and be tailored to target certain types of cancer, such as breast or colon cancer, specifically. “This means that we can investigate thousands of genes at a time which will allow us to comb through the human genome with previously unattainable efficiency,” said Dr. Hannon. “We have already identified several genes that appear to be selectively required by certain breast cancer cells but not by normal cells.”
As this new technique is used to discover more and more cancer proliferation and survival genes associated with different kinds of cancer, it could introduce a new era in cancer treatment. “We have opened a door into a whole new world of cancer treatment possibilities,” Dr. Hannon said.
Written by: Communications Department | publicaffairs@cshl.edu | 516-367-8455
Citation
“Cancer Proliferation Gene Discovery Through Functional Genomics” appeared in Science on February 1, 2008. The complete citation is as follows: Michael R. Schlabach, Ji Luo, Nicole L. Solimini, Guang Hu, Qikai Xu, Mamie Z. Li, Zhenming Zhao, Agata Smogorzewska, Matthew E. Sowa, Xiaolu L. Ang, Thomas F. Westbrook, Anthony C. Liang, Kenneth Chang, Jennifer A. Hackett, J. Wade Harper, Gregory J. Hannon, Stephen J. Elledge. The paper is available online here.
“Profiling Essential Genes in Human Mammary Cells by Multiplex RNAi Screening” appears in the same issue of Science and is available online here. Its full citation is as follows: Jose M. Silva, Krista Marran, Joel S. Parker, Javier Silva, Michael Golding, Michael R. Schlabach, Stephen J. Elledge, Gregory J. Hannon, Kenneth Chang.