Cold Spring Harbor Laboratory (CSHL) Assistant Professor Peter Koo and collaborator Matt Ploenzke reported a way to train machines to predict the function of DNA sequences. They used “neural nets,” a type of artificial intelligence (AI) typically used to classify images. Teaching the neural net to predict the function of short stretches of DNA allowed it to work up to deciphering larger patterns. The researchers hope to analyze more complex DNA sequences that regulate gene activity critical to development and disease.
Machine-learning researchers can train a brain-like “neural net” computer to recognize objects, such as cats or airplanes, by showing it many images of each. Testing the success of training requires showing the machine a new picture of a cat or an airplane and seeing if it classifies it correctly. But, when researchers apply this technology to analyzing DNA patterns, they have a problem. Humans can’t recognize the patterns, so they may not be able to tell if the computer identifies the right thing. Neural nets learn and make decisions independently of their human programmers. Researchers refer to this hidden process as a “black box.” It is hard to trust the machine’s outputs if we don’t know what is happening in the box.
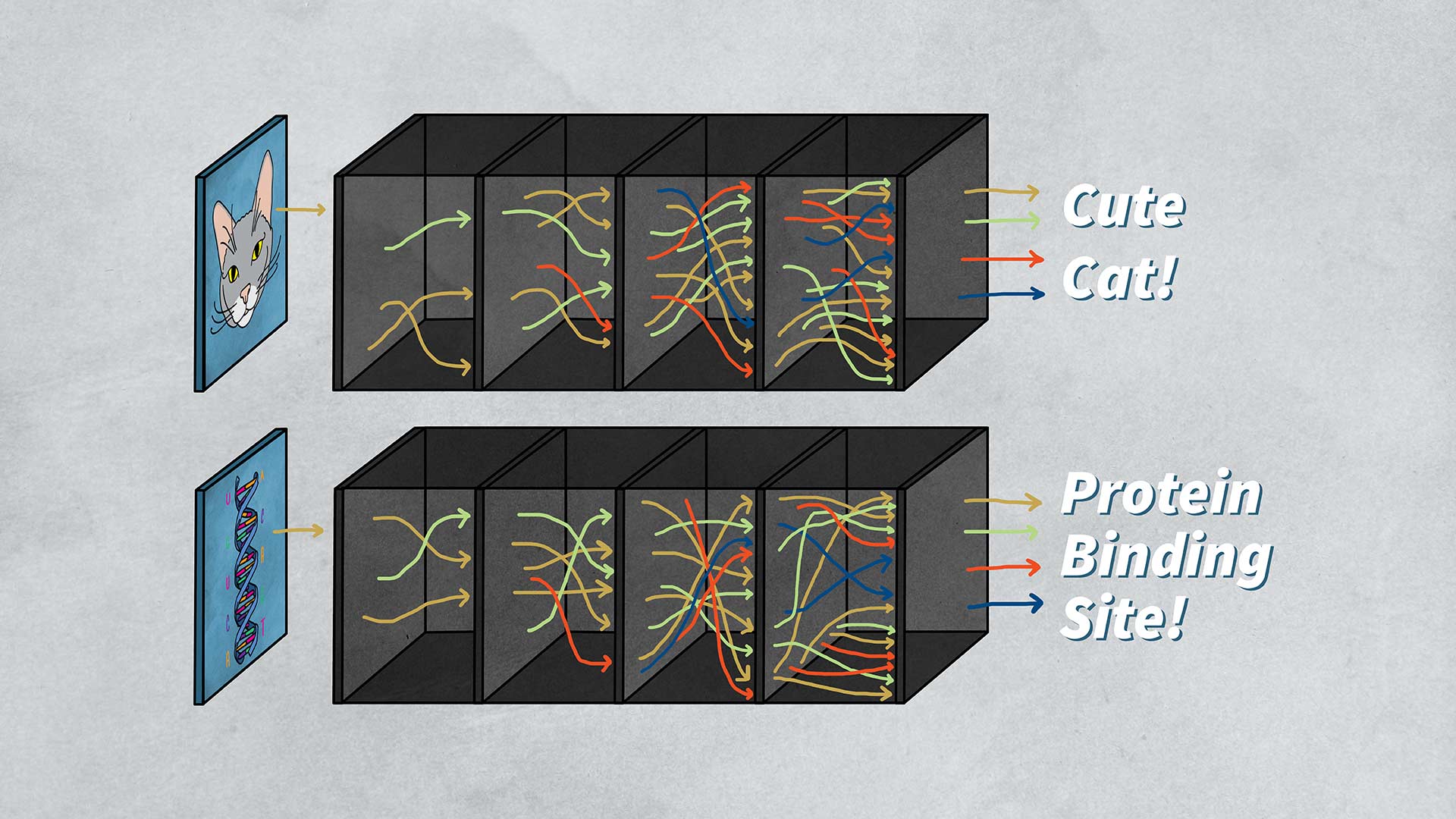
Koo and his team fed DNA (genomic) sequences into a specific kind of neural network called a convolutional neural network (CNN), which resembles how animal brains process images. Koo says:
“It can be quite easy to interpret these neural networks because they’ll just point to, let’s say, whiskers of a cat. And so that’s why it’s a cat versus an airplane. In genomics, it’s not so straightforward because genomic sequences aren’t in a form where humans really understand any of the patterns that these neural networks point to.”
Koo’s research, reported in the journal Nature Machine Intelligence, introduced a new method to teach important DNA patterns to one layer of his CNN. This allowed his neural network to build on the data to identify more complex patterns. Koo’s discovery makes it possible to peek inside the black box and identify some key features that lead to the computer’s decision-making process.
But Koo has a larger purpose in mind for the field of artificial intelligence. There are two ways to improve a neural net: interpretability and robustness. Interpretability refers to the ability of humans to decipher why machines give a certain prediction. The ability to produce an answer even with mistakes in the data is called robustness. Usually, researchers focus on one or the other. Koo says:
“What my research is trying to do is bridge these two together because I don’t think they’re separate entities. I think that we get better interpretability if our models are more robust.”
Koo hopes that if a machine can find robust and interpretable DNA patterns related to gene regulation, it will help geneticists understand how mutations affect cancer and other diseases.
Written by: Jasmine Lee, Content Developer/Communicator | publicaffairs@cshl.edu | 516-367-8845
Funding
National Cancer Institute, Simons Center for Quantitative Biology, National Institutes of Health
Citation
Koo, P., et al., “Improving representations of genomic sequence motifs in convolutional networks with exponential activations”, Nature Machine Intelligence, February 8, 2021. DOI: https://doi.org/10.1101/2020.06.14.150706
Principal Investigator

Peter Koo
Associate Professor
Cancer Center Member
Ph.D., Yale University, 2015