Cold Spring Harbor, NY — Most people understand genes to be specific segments of DNA that determine traits or diseases that are inherited. Textbooks suggest that genes are copied (“transcribed”) into RNA molecules, which are then used as templates for making protein—the highly diverse set of molecules that act as building blocks and engines of our cells. The truth, it now appears, is not so simple.
As part of a huge collaborative effort called ENCODE (Encyclopedia of DNA Elements), a research team led by Cold Spring Harbor Laboratory (CSHL) Professor Thomas Gingeras, Ph.D., today publishes a genome-wide analysis of RNA messages, called transcripts, produced within human cells.
Their analysis—one component of a massive release of research results by ENCODE teams from 32 institutes in 5 countries, with 30 papers appearing today in 3 different high-level scientific journals—shows that three-quarters of the genome is capable of being transcribed. This figure is important because it indicates that nearly all of our genome is dynamic and active. It stands in marked contrast to consensus views prior to ENCODE’s comprehensive research efforts, which suggested that only the small protein-encoding fraction of the genome was transcribed, and therefore important.
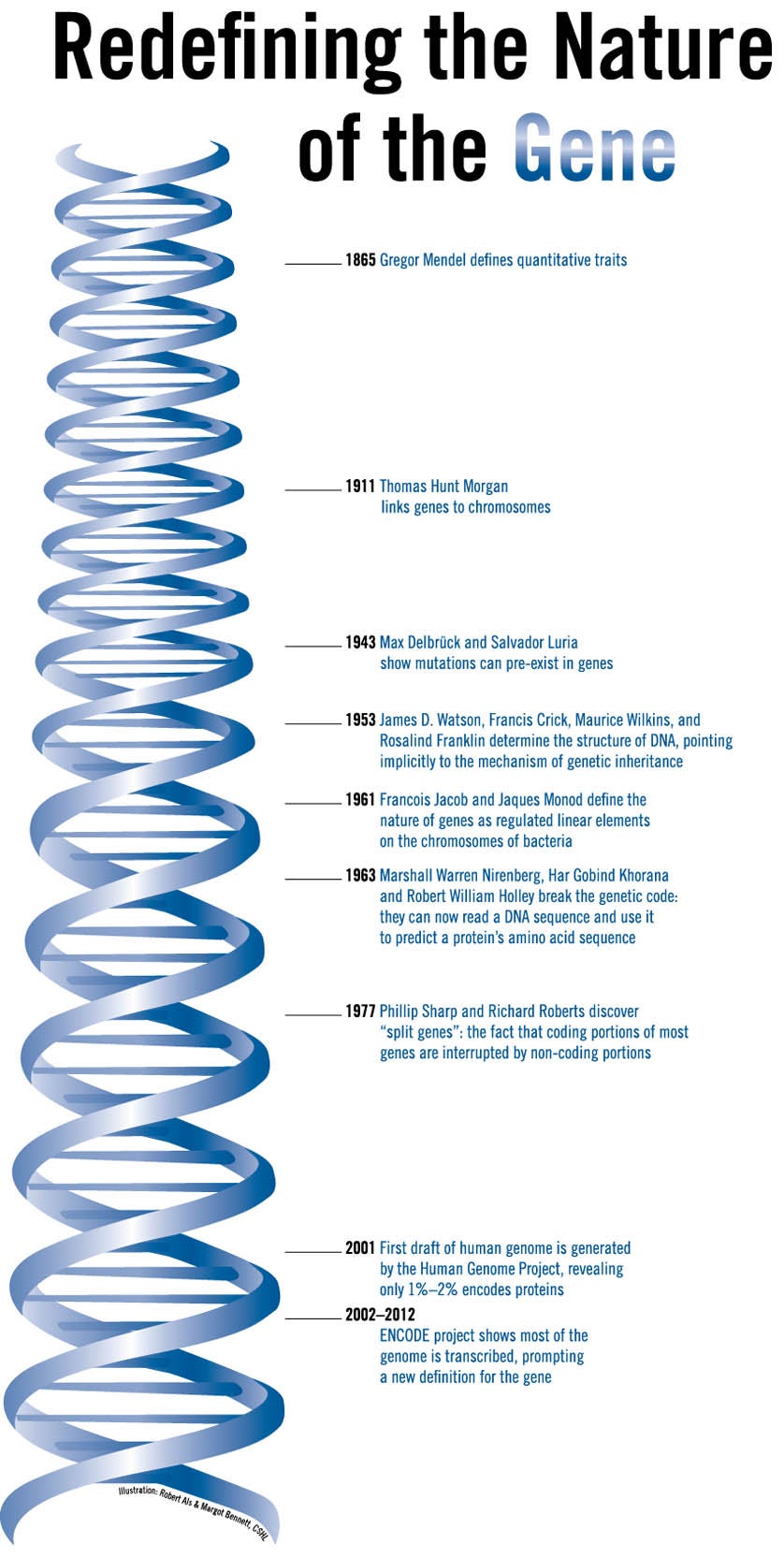
Credit: Margot Bennett and Robert Als.
The vast amount of data generated with advanced technologies by Gingeras’ group and others in the ENCODE project is likely to radically change the prevailing understanding of what defines a gene, the unit we routinely use, for instance, to speak of inheritable traits like eye color or to explain the causes of and susceptibility to most diseases, running the gamut from cancer to schizophrenia to heart disease.
In 2003 the ENCODE project consortium was set up by the U.S. government’s National Human Genome Research Institute (NHGRI) to examine the newly minted sequence of the human genome in greater depth. At the time, the genome was thought of as a linear molecule of DNA with “genes” being contained within isolated sections that make up just 1%-2% of its total length. The long stretches of DNA between these gene islands were once thought to be mostly functionless spacers, padding, or even “junk DNA.”
Through the work of Gingeras and others in this latest phase of the ENCODE project consortium, we now know that most of the DNA around protein-encoding genes is also capable of being transcribed into RNA—another way of saying that it has the potential of performing useful functions in cells.
In preliminary ENCODE results published in 2007, the researchers closely examined about 1% of the human genome. The initial results showed that much more of our DNA could be transcribed than previously thought. Far from being padding, many of these RNA messages appeared to be functional.
The Gingeras lab discovered potentially new classes of functional RNAs in this preliminary work. The additional knowledge that parts of one gene or functional RNA can reside within another were surprising, and suggested a picture of the architecture of our genome that was much more complex than previously thought.
What the new ENCODE data reveals
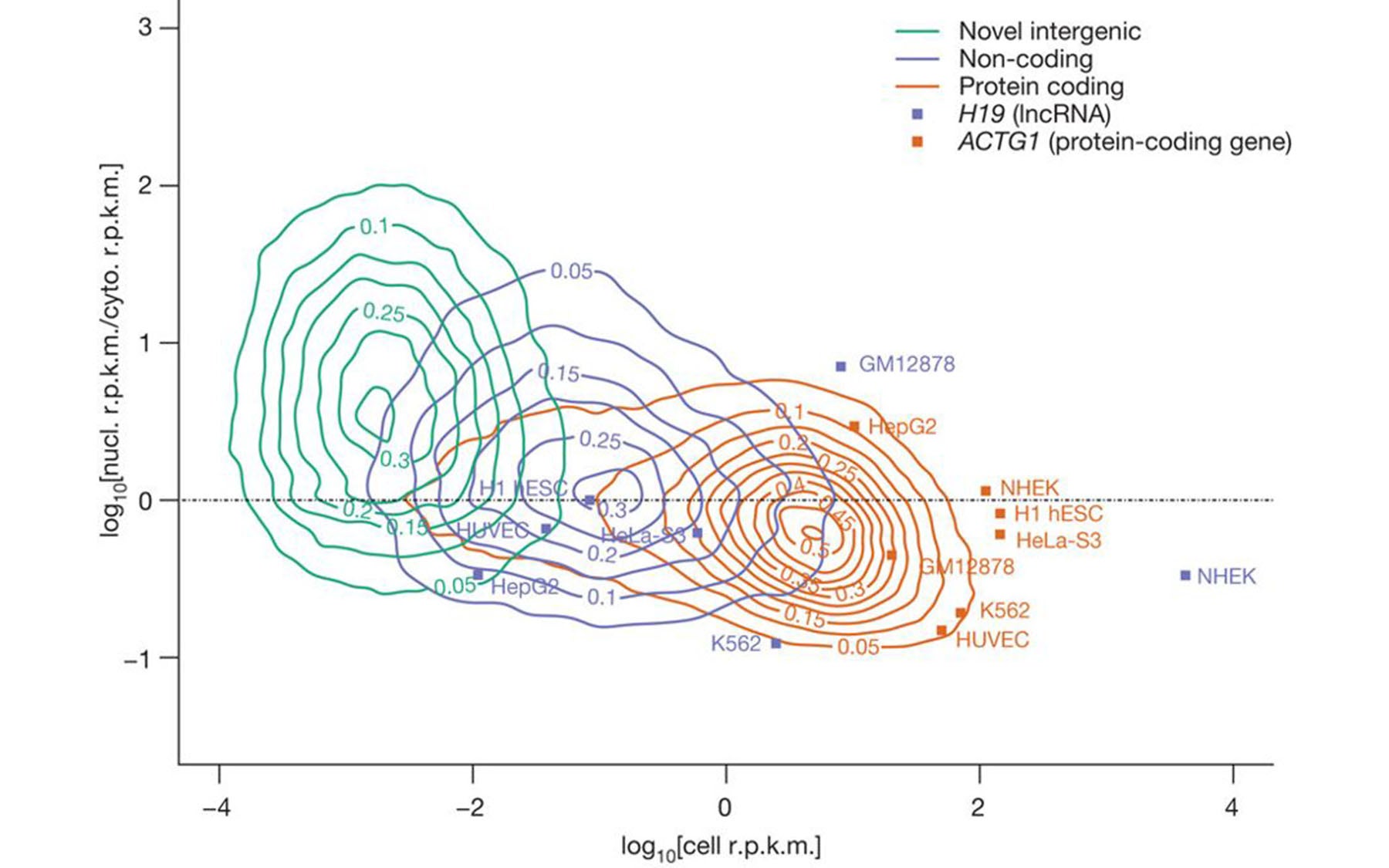
Two of the 30 papers published by Gingeras and other ENCODE colleagues, including CSHL Professor and HHMI Investigator Gregory Hannon, Ph.D., who is also a co-author in this study, today mark the culmination of project’s second phase. What distinguishes the data analyzed in this phase is comprehensiveness. The initial observations of 2007 are now extended to cover the entire human genome—a tour-de-force effort in which the transcribed RNA from different sub-cellular compartments of 15 human cell lines was analyzed.
Although the results vary between cell lines, a consensus picture is emerging. In addition to showing that up to three-quarters of our DNA may be transcribed into RNA, the data strongly suggests, according to Gingeras, that a large percent of non-protein-coding RNAs are localized within cells in a manner consistent with their having functional roles.
The current outstanding question concerns the nature and range of those functions. It is thought that these “non-coding” RNA transcripts act something like components of a giant, complex switchboard, controlling a network of many events in the cell by regulating the processes of replication, transcription and translation—that is, the copying of DNA and the making of proteins based on information carried by messenger RNAs.
With the understanding that so much of our DNA can be transcribed into RNA comes the realization that there is much less space between what we previously thought of as genes, Gingeras points out.
“We see the boundaries of what were assumed to be the regions between genes shrinking in length,” he says, “and genic regions making many overlapping RNAs.” It appears, he continues, that the boundaries of conventionally described genes are melding together, challenging the notion that a gene is a discrete, localized region of a genome separated by inert DNA. “New definitions of a gene are needed,” Gingeras says.
What are the practical implications? According to Gingeras, they include being able to identify possible causes for natural traits such as height or hair loss and disease states such as cancer. Many genetic variations associated with a trait often map to what were formally believed to be “spacer” regions.
“With our increasingly deeper understanding that such regions are related to the neighboring or “distal” protein coding regions—via the creation of non-coding RNAs—we will now seek underlying explanations of the association of the genetic variation and traits of interest.” This topic is explored in a second paper published today that summarizes the finding of all the consortium groups participating in the current phase of the ENCODE project: The ENCODE Project Consortium. 2012. An integrated encyclopedia of DNA elements in the human genome. Nature: doi:10.1038/nature11247.
“Exploration of the genome is akin to our efforts at exploring our physical universe,” Gingeras says. “We expect to be amazed and excited by our future efforts to map and explore our personal genetic universes.”
Written by: Edward Brydon, Science Writer | publicaffairs@cshl.edu | 516-367-8455
Funding
The research described in this release was supported by the National Human Genome Research Institute (NHGRI) production grants U54HG004557, U54HG004555, U54HG004576 and U54HG004558, and by the NHGRI pilot project grant R01HG003700. It was also supported by the NHGRI ARRA stimulus grant 1RC2HG005591, the National Science Foundation (NSF) grant 127375, the European Research Council (ERC) grant 249968, a research grant for the RIKEN Omics Science Center from the Japanese Ministry of Education, Culture, Sports, Science and Technology, and grants BIO2011-26205, CSD2007-00050 and INB-GNV1 from the Spanish Ministry of Science.
Citation
“Landscape of transcription in human cells” is published online in Nature on September 5, 2012. The authors are: Sarah Djebali, Carrie A. Davis and 83 others. The paper can be obtained online at doi:10.1038/nature11233. Other new ENCODE results can be found in the following journals: Nature (6 papers); Genome Research (18 papers); and Genome Biology (6 papers).
The full ENCODE Consortium data sets can be freely accessed through the ENCODE project portal as well as at the University of California at Santa Cruz genome browser, the National Center for Biotechnology Information, and the European Bioinformatics Institute. Topic threads that run through several different papers can be explored via the ENCODE microsite page at Nature.com/encode.
Principal Investigator

Thomas Gingeras
Professor
Cancer Center Member
Ph.D., New York University, 1976